Compound details
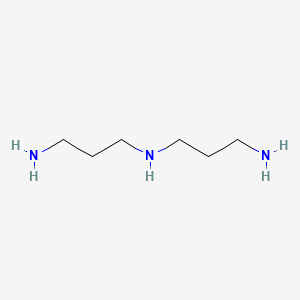
Norspermidine
| Compound ID | CDAMM01846 |
|---|---|
| Common name | Norspermidine | IUPAC name | N\'-(3-aminopropyl)propane-1,3-diamine |
| Molecular formula | C6H17N3 |
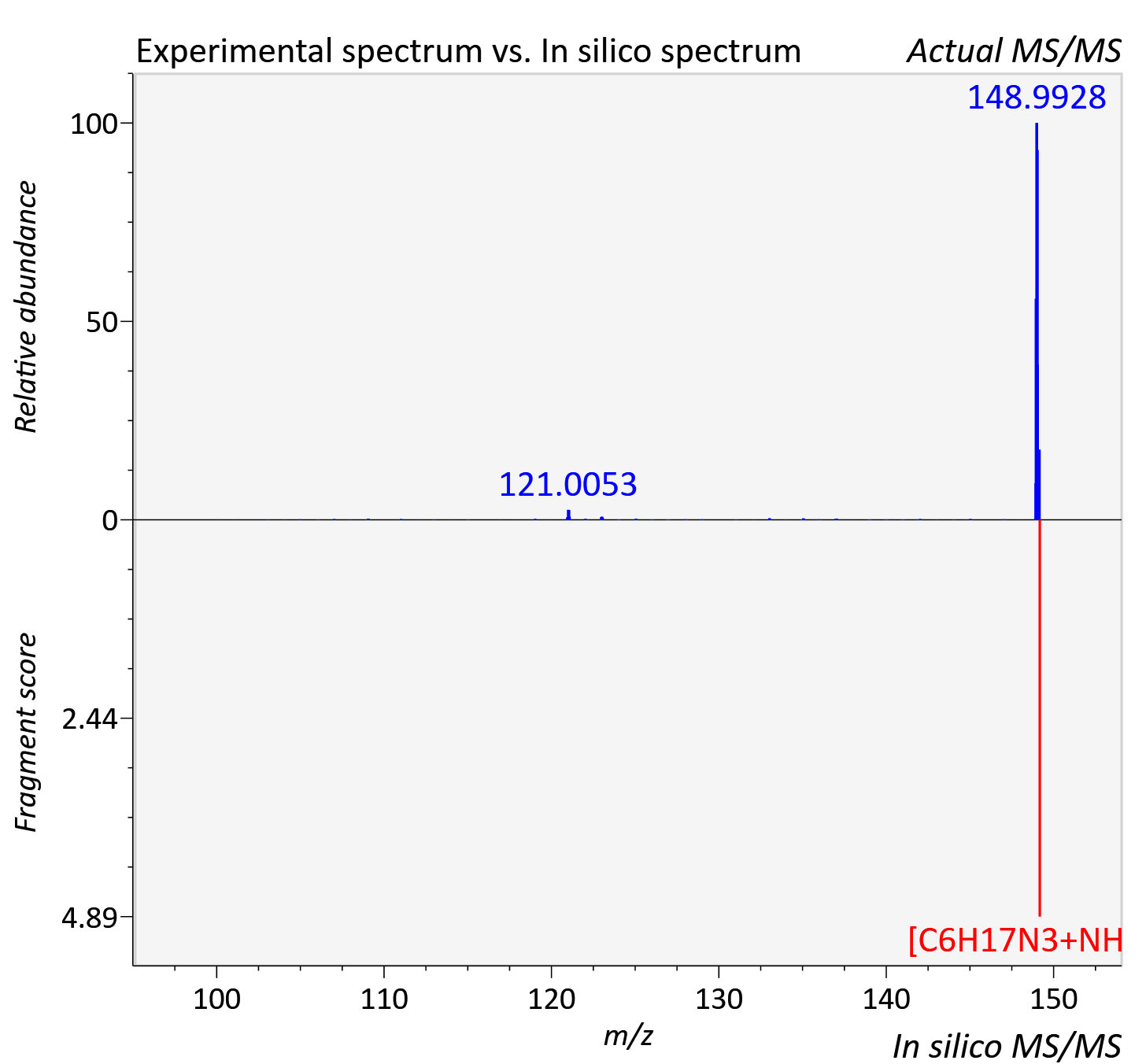
Experimental data
| Retention time | 14.7 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 149.176 | Theoretical mz | 149.176 |
| Error | 2.17 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.255 |
Identifiers and class information
| Inchi key | OTBHHUPVCYLGQO-UHFFFAOYSA-N |
|---|---|
| Smiles | NCCCNCCCN |
| Superclass | Organic nitrogen compounds |
| Class | Organonitrogen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| P48066 | SLC6A11 | GABA transporter 3 | T38200 | SEA |
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| Q05586 | GRIN1 | Glutamate NMDA receptor; GRIN1/GRIN2B | T62276 | SEA |
| Q12879 | GRIN2A | Glutamate NMDA receptor; GRIN1/GRIN2A | T22285 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| Q6QHF9 | PAOX | Polyamine oxidase | T88699 | SEA |
| P42575 | CASP2 | Caspase-2 | T09128 | SEA |
| Q13224 | GRIN2B | Glutamate [NMDA] receptor subunit epsilon 2 | T76414 | SEA |
| Q9UN88 | GABRQ | GABA(A) receptor theta | T15161 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T38200 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P48066 | SLC6A11 |
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T62276 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q05586 | GRIN1 |
| T62276 | DI0182 | HIV-infected patients with tuberculosis | [ICD-11: 1B10-1B14] | Q05586 | GRIN1 |
| T22285 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | Q12879 | GRIN2A |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T76414 | DI0024 | Alpha-1-antitrypsin deficiency | [ICD-11: 5C5A] | Q13224 | GRIN2B |
