Compound details
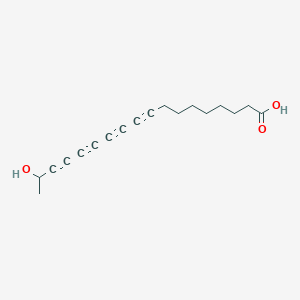
(S)-17-Hydroxy-9,11,13,15-octadecatetraynoic acid
| Compound ID | CDAMM01845 |
|---|---|
| Common name | (S)-17-Hydroxy-9,11,13,15-octadecatetraynoic acid | IUPAC name | 17-hydroxyoctadeca-9,11,13,15-tetraynoic acid |
| Molecular formula | C18H20O3 |
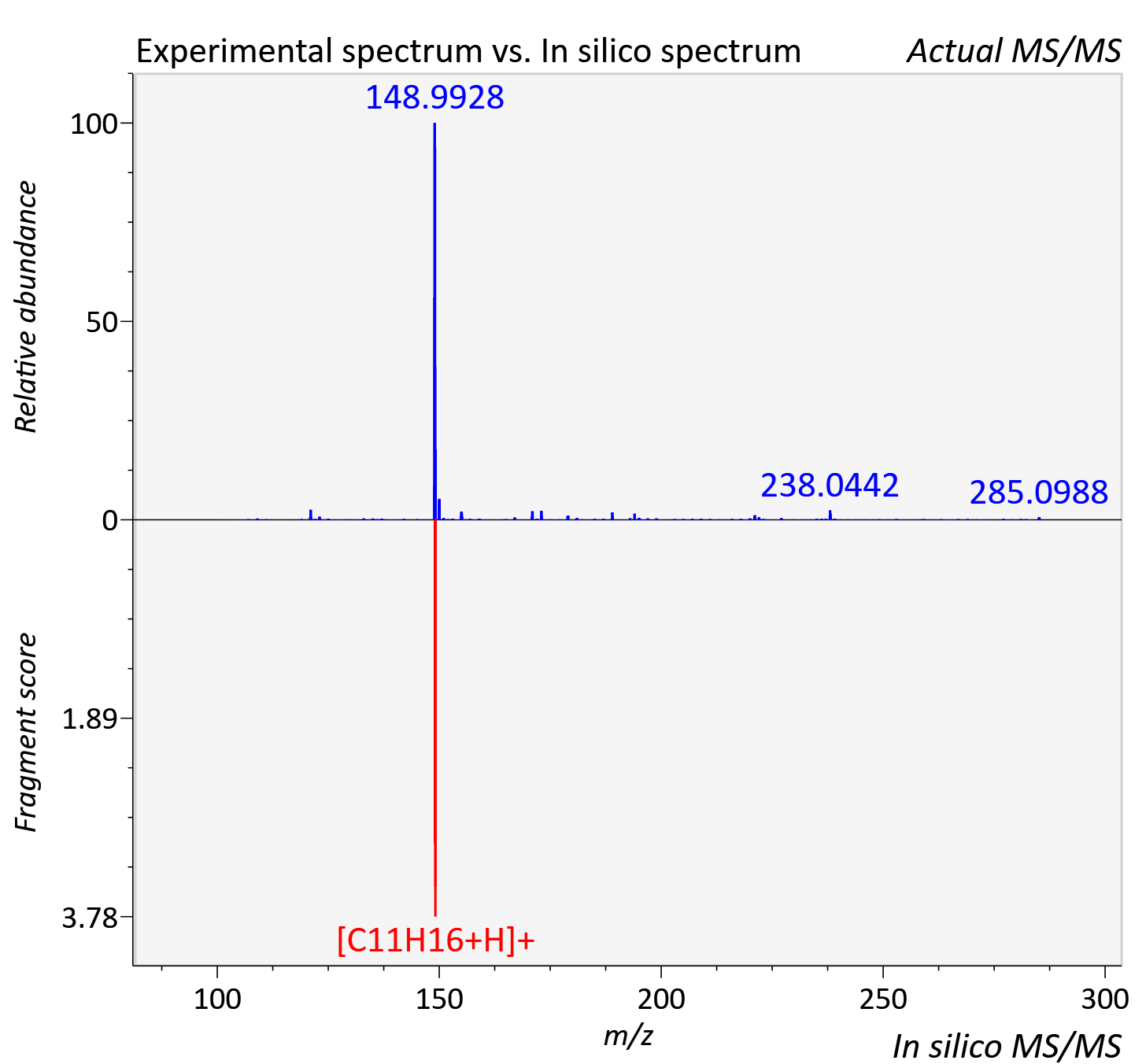
Experimental data
| Retention time | 14.7 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 285.147 | Theoretical mz | 285.148 |
| Error | 3.07 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.055 |
Identifiers and class information
| Inchi key | MTWGWIOCIREVRF-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(O)CCCCCCCC#CC#CC#CC#CC(O)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| P48066 | SLC6A11 | GABA transporter 3 | T38200 | SEA |
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| Q07075 | ENPEP | Aminopeptidase A | T31956 | SEA |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| Q9UN88 | GABRQ | GABA(A) receptor theta | T15161 | SEA |
| P78329 | CYP4F2 | Cytochrome P450 4f | T82604 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T38200 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P48066 | SLC6A11 |
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T31956 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q07075 | ENPEP |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
