Compound details
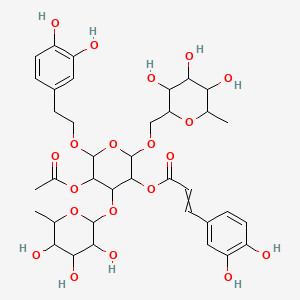
Brandioside
| Compound ID | CDAMM01831 |
|---|---|
| Common name | Brandioside | IUPAC name | [5-acetyloxy-6-[2-(3,4-dihydroxyphenyl)ethoxy]-2-[(3,4,5-trihydroxy-6-methyloxan-2-yl)methoxy]-4-(3,4,5-trihydroxy-6-methyloxan-2-yl)oxyoxan-3-yl] 3-(3,4-dihydroxyphenyl)prop-2-enoate |
| Molecular formula | C37H48O20 |
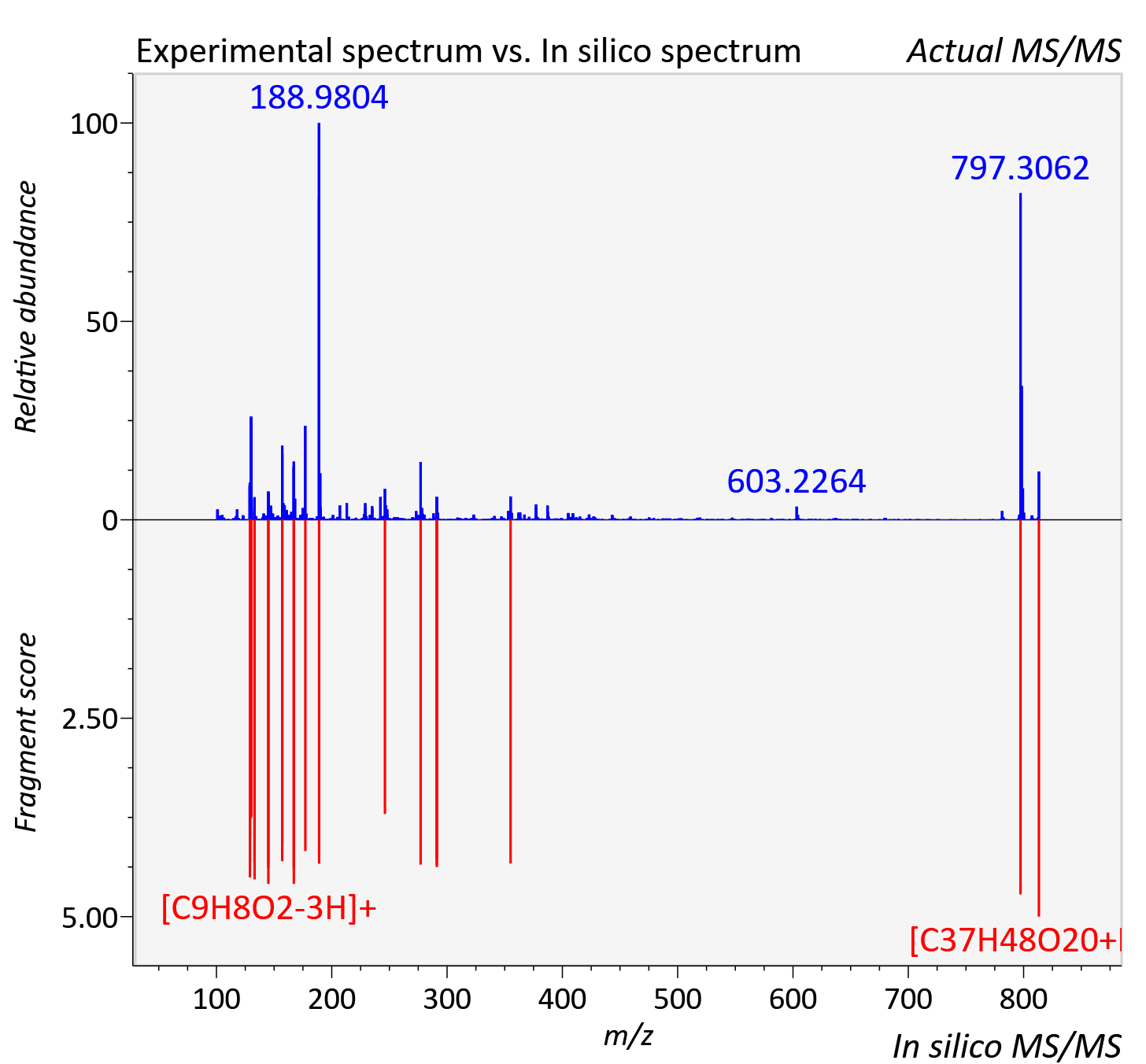
Experimental data
| Retention time | 7.79 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 813.281 | Theoretical mz | 813.281 |
| Error | 0.46 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.9545 |
Identifiers and class information
| Inchi key | SAUVJZCYCNLQSI-RMKNXTFCNA-N |
|---|---|
| Smiles | O=C(OC1C(OCC2OC(C)C(O)C(O)C2O)OC(OCCC3=CC=C(O)C(O)=C3)C(OC(=O)C)C1OC4OC(C)C(O)C(O)C4O)C=CC5=CC=C(O)C(O)=C5 |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SEA |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SEA |
| P02766 | TTR | Transthyretin | T86462 | SEA |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P08253 | MMP2 | Matrix metalloproteinase 2 | T68251 | SwissTargetPrediction |
| P39900 | MMP12 | Matrix metalloproteinase 12 | T03500 | SwissTargetPrediction |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| P04746 | AMY2A | Pancreatic alpha-amylase | T86918 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T86462 | DI0026 | Amyloidosis | [ICD-11: 5D00] | P02766 | TTR |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T68251 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08253 | MMP2 |
| T03500 | DI0037 | Asthma | [ICD-11: CA23] | P39900 | MMP12 |
| T03500 | DI0238 | Lung cancer | [ICD-11: 2C25] | P39900 | MMP12 |
| T03500 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P39900 | MMP12 |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T86918 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P04746 | AMY2A |
| T86918 | DI0328 | Pancreatic malfunction | [ICD-11: DC30-DC3Z] | P04746 | AMY2A |
