Compound details
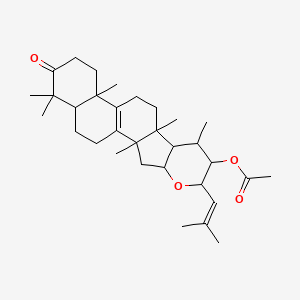
Echinodone
| Compound ID | CDAMM01800 |
|---|---|
| Common name | Echinodone | IUPAC name | [2,8,10,14,18,18-hexamethyl-6-(2-methylprop-1-enyl)-17-oxo-5-oxapentacyclo[11.8.0.02,10.04,9.014,19]henicos-1(13)-en-7-yl] acetate |
| Molecular formula | C32H48O4 |
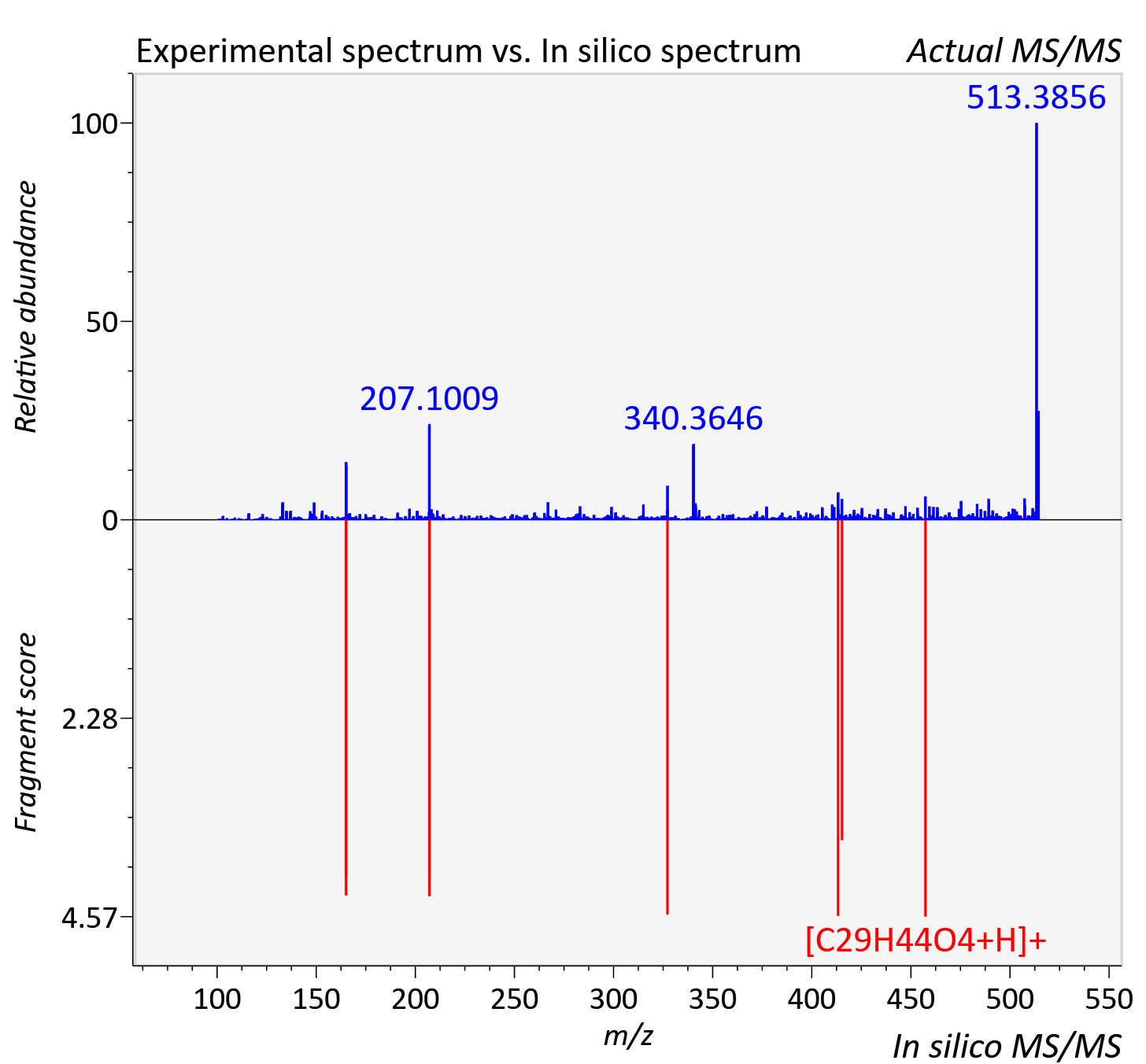
Experimental data
| Retention time | 15.6 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 514.39 | Theoretical mz | 514.389 |
| Error | 1.32 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.584 |
Identifiers and class information
| Inchi key | ACPQHJVBMJBRJO-BZZCOUAPNA-N |
|---|---|
| Smiles | O=C(OC1C(OC2CC3(C4=C(CCC3(C)C2C1C)C5(C)CCC(=O)C(C)(C)C5CC4)C)C=C(C)C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SwissTargetPrediction |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SwissTargetPrediction |
| P05771 | PRKCB | Protein kinase C beta | T40276 | SwissTargetPrediction |
| P50579 | METAP2 | Methionine aminopeptidase 2 | T75596 | SwissTargetPrediction |
| P25024 | CXCR1 | Interleukin-8 receptor A | T00884 | SwissTargetPrediction |
| P55085 | F2RL1 | Proteinase-activated receptor 2 | T62841 | SwissTargetPrediction |
| P01584 | IL1B | Interleukin-1 beta | T42000 | SwissTargetPrediction |
| Q13526 | PIN1 | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | T16308 | SEA |
| P05129 | PRKCG | Protein kinase C gamma | T47107 | SwissTargetPrediction |
| P41252 | IARS | Isoleucyl-tRNA synthetase | T46515 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T40276 | DI0123 | Diffuse large B-cell lymphoma | [ICD-11: 2A81] | P05771 | PRKCB |
| T40276 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P05771 | PRKCB |
| T40276 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P05771 | PRKCB |
| T75596 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P50579 | METAP2 |
| T00884 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P25024 | CXCR1 |
| T62841 | DI0087 | Chronic pain | [ICD-11: MG30] | P55085 | F2RL1 |
| T42000 | DI0267 | Mineral excesses | [ICD-11: 5B91] | P01584 | IL1B |
| T42000 | DI0269 | Monogenic autoinflammatory syndrome | [ICD-11: 4A60] | P01584 | IL1B |
| T42000 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P01584 | IL1B |
| T42000 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P01584 | IL1B |
| T16308 | DI0238 | Lung cancer | [ICD-11: 2C25] | Q13526 | PIN1 |
| T47107 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P05129 | PRKCG |
| T47107 | DI0248 | Mastocytosis | [ICD-11: 2A21] | P05129 | PRKCG |
