Compound details
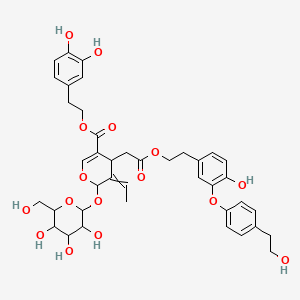
Insuloside
| Compound ID | CDAMM01796 |
|---|---|
| Common name | Insuloside | IUPAC name | 2-(3,4-dihydroxyphenyl)ethyl 5-ethylidene-4-[2-[2-[4-hydroxy-3-[4-(2-hydroxyethyl)phenoxy]phenyl]ethoxy]-2-oxoethyl]-6-[3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy-4H-pyran-3-carboxylate |
| Molecular formula | C40H46O16 |
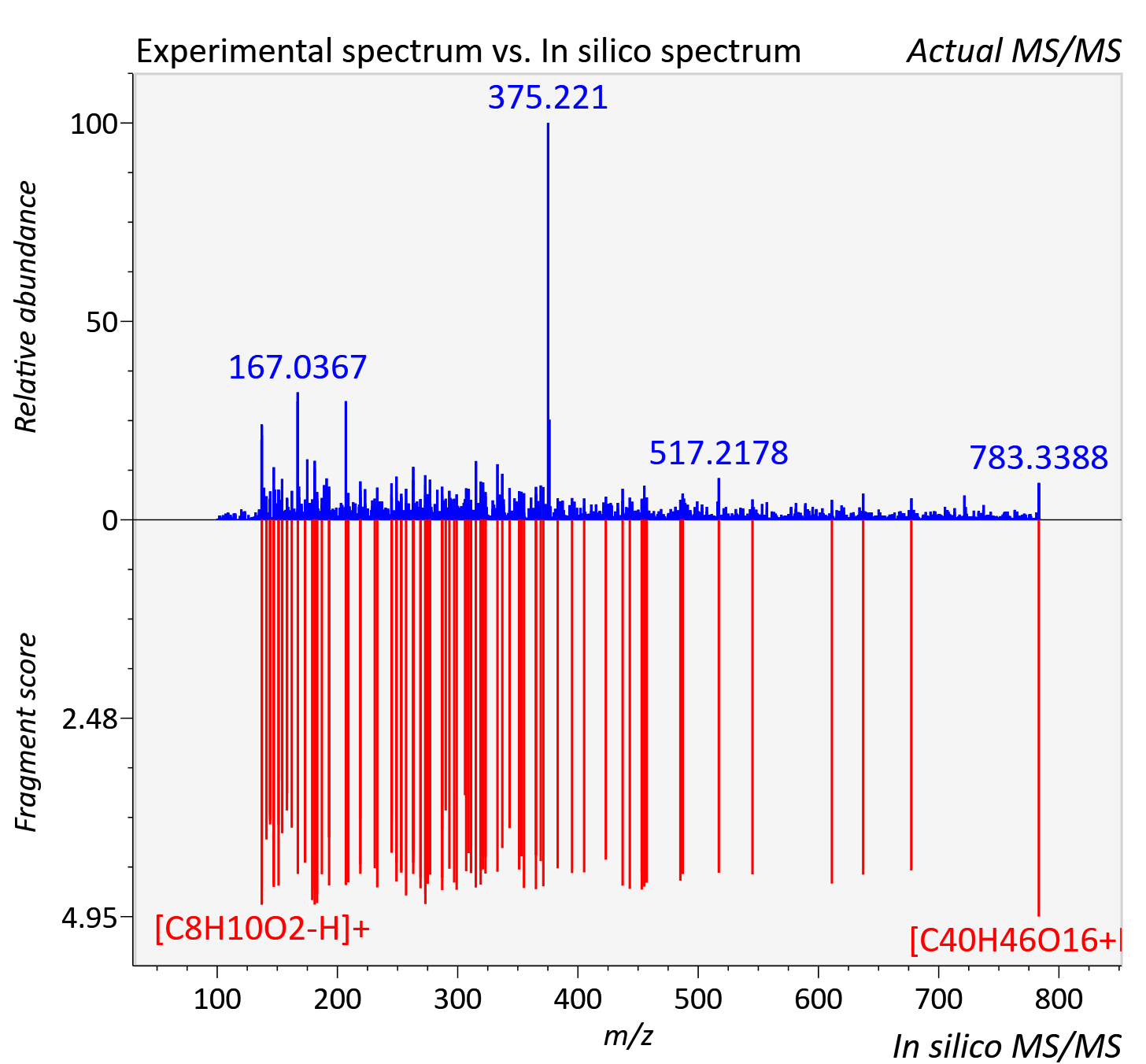
Experimental data
| Retention time | 6.01 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 783.293 | Theoretical mz | 783.286 |
| Error | 8.71 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0613 |
Identifiers and class information
| Inchi key | SREDLZCCQWSQLN-QBQOLSIMNA-N |
|---|---|
| Smiles | O=C(OCCC1=CC=C(O)C(O)=C1)C2=COC(OC3OC(CO)C(O)C(O)C3O)C(=CC)C2CC(=O)OCCC4=CC=C(O)C(OC5=CC=C(C=C5)CCO)=C4 |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P01112 | HRAS | Transforming protein p21/H-Ras-1 | T71081 | SEA |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| Q9NUW8 | TDP1 | Tyrosyl-DNA phosphodiesterase 1 | T33492 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
