Compound details
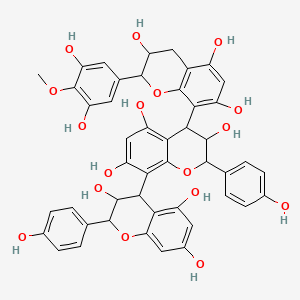
[Epiafzelechin-(4beta->8)]2-4\'-O-methyl-epigallocatechin
| Compound ID | CDAMM01790 |
|---|---|
| Common name | [Epiafzelechin-(4beta->8)]2-4\'-O-methyl-epigallocatechin | IUPAC name | 4-[2-(3,5-dihydroxy-4-methoxyphenyl)-3,5,7-trihydroxy-3,4-dihydro-2H-chromen-8-yl]-2-(4-hydroxyphenyl)-8-[3,5,7-trihydroxy-2-(4-hydroxyphenyl)-3,4-dihydro-2H-chromen-4-yl]-3,4-dihydro-2H-chromene-3,5,7-triol |
| Molecular formula | C46H40O17 |
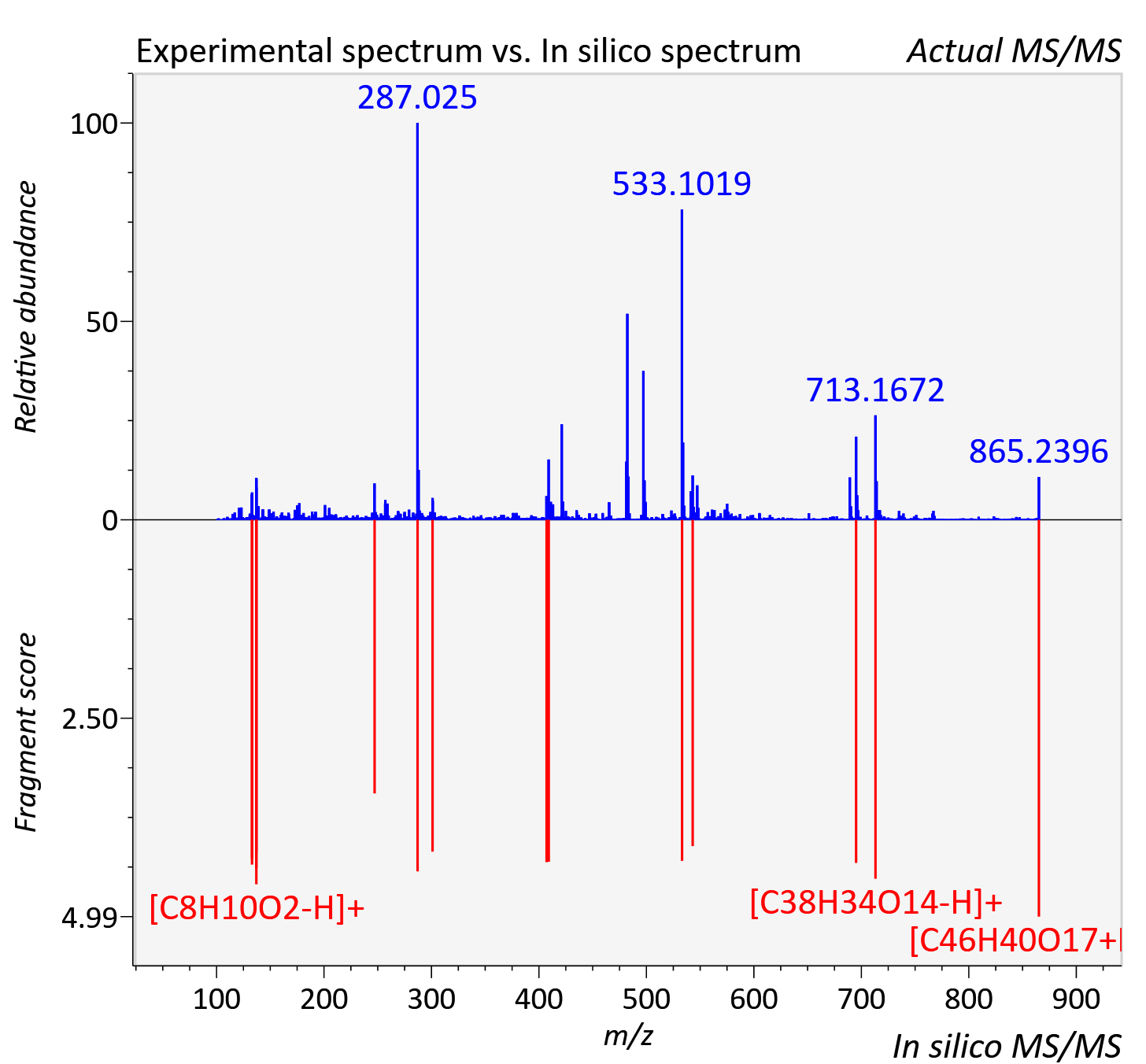
Experimental data
| Retention time | 3.2 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 865.235 | Theoretical mz | 865.234 |
| Error | 1.18 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.4916 |
Identifiers and class information
| Inchi key | QKCQZHCKCYRFJX-UZWUUNAJNA-N |
|---|---|
| Smiles | OC1=CC=C(C=C1)C2OC=3C=C(O)C=C(O)C3C(C4=C(O)C=C(O)C5=C4OC(C6=CC=C(O)C=C6)C(O)C5C=7C(O)=CC(O)=C8C7OC(C9=CC(O)=C(OC)C(O)=C9)C(O)C8)C2O |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| P08253 | MMP2 | Matrix metalloproteinase 2 | T68251 | SwissTargetPrediction |
| P14780 | MMP9 | Matrix metalloproteinase 9 | T54156 | SwissTargetPrediction |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| P20151 | KLK2 | Kallikrein 2 | T01908 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T68251 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08253 | MMP2 |
| T54156 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P14780 | MMP9 |
| T54156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P14780 | MMP9 |
| T54156 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P14780 | MMP9 |
| T54156 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P14780 | MMP9 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T01908 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P20151 | KLK2 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
