Compound details
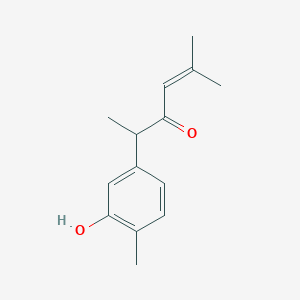
2-(3-Hydroxy-4-methylphenyl)-5-methyl-4-hexen-3-one
| Compound ID | CDAMM01748 |
|---|---|
| Common name | 2-(3-Hydroxy-4-methylphenyl)-5-methyl-4-hexen-3-one | IUPAC name | 2-(3-hydroxy-4-methylphenyl)-5-methylhex-4-en-3-one |
| Molecular formula | C14H18O2 |
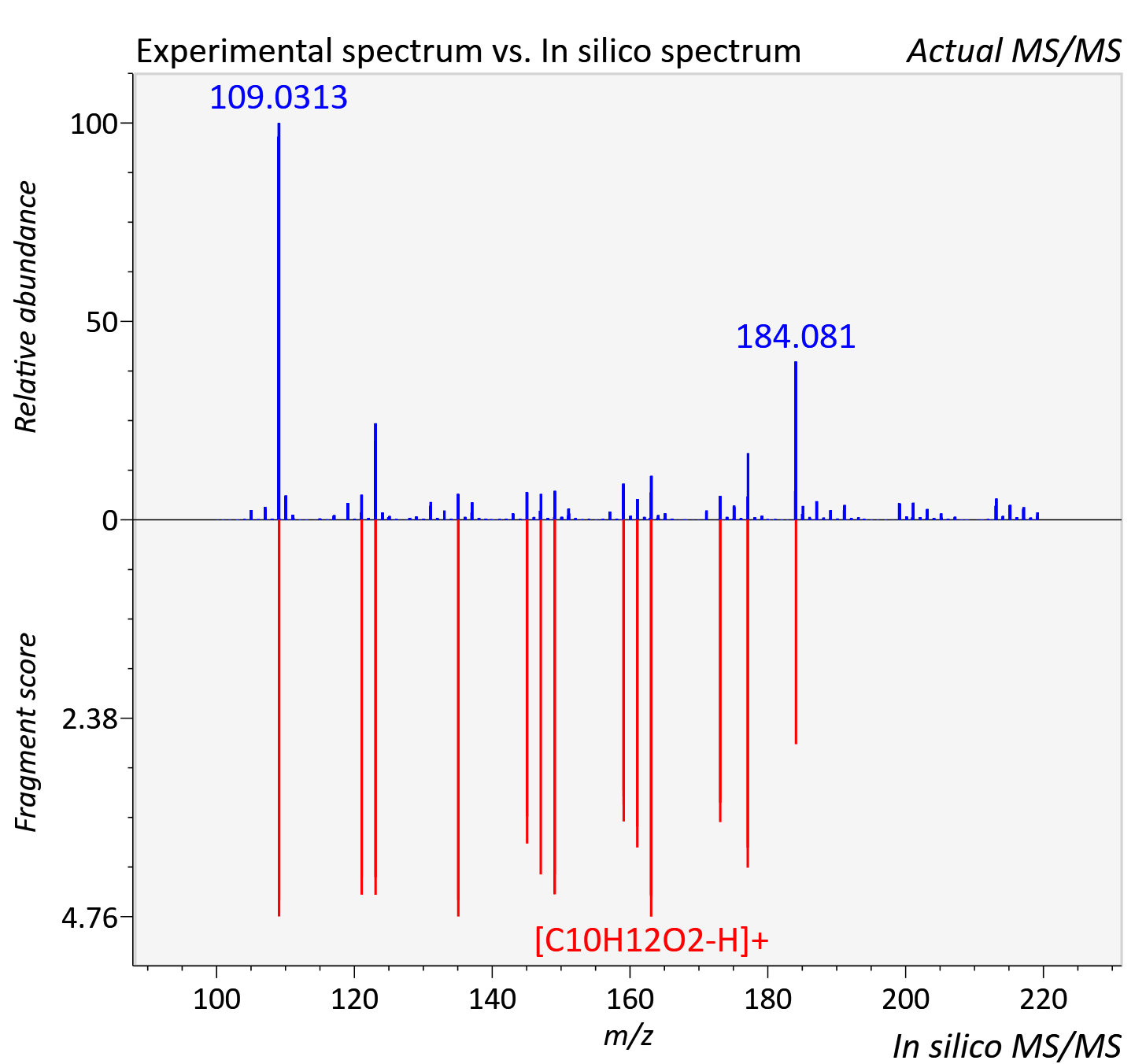
Experimental data
| Retention time | 19 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 219.139 | Theoretical mz | 219.138 |
| Error | 5.13 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.3924 |
Identifiers and class information
| Inchi key | WZHLEYQOLDOTEY-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(C=C(C)C)C(C1=CC=C(C(O)=C1)C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T22658 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P10145 | CXCL8 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
