Compound details
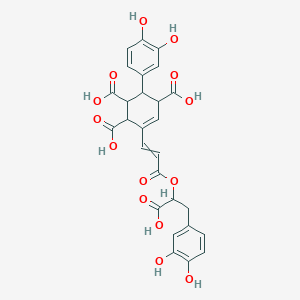
Yunnaneic acid E
| Compound ID | CDAMM01721 |
|---|---|
| Common name | Yunnaneic acid E | IUPAC name | 6-[3-[1-carboxy-2-(3,4-dihydroxyphenyl)ethoxy]-3-oxoprop-1-enyl]-3-(3,4-dihydroxyphenyl)cyclohex-5-ene-1,2,4-tricarboxylic acid |
| Molecular formula | C27H24O14 |
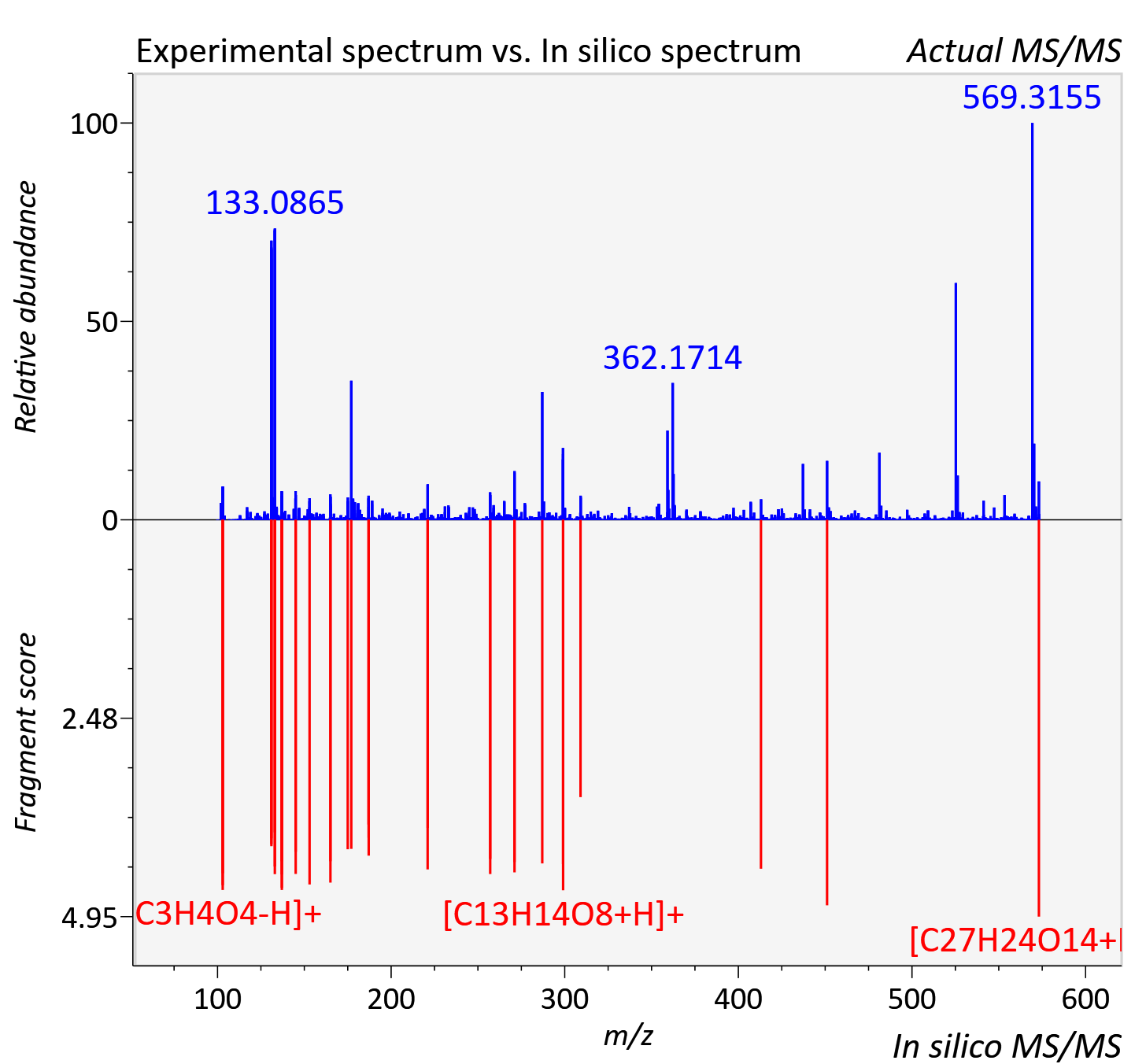
Experimental data
| Retention time | 3.99 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 573.116 | Theoretical mz | 573.124 |
| Error | 13.92 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.6009 |
Identifiers and class information
| Inchi key | JSOPGXFFNOKRAX-FUAGRLKYNA-N |
|---|---|
| Smiles | O=C(OC(C(=O)O)CC1=CC=C(O)C(O)=C1)C=CC2=CC(C(=O)O)C(C3=CC=C(O)C(O)=C3)C(C(=O)O)C2C(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
