Compound details
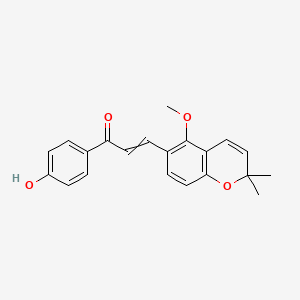
Licoagrochalcone B
| Compound ID | CDAMM01703 |
|---|---|
| Common name | Licoagrochalcone B | IUPAC name | 1-(4-hydroxyphenyl)-3-(5-methoxy-2,2-dimethylchromen-6-yl)prop-2-en-1-one |
| Molecular formula | C21H20O4 |
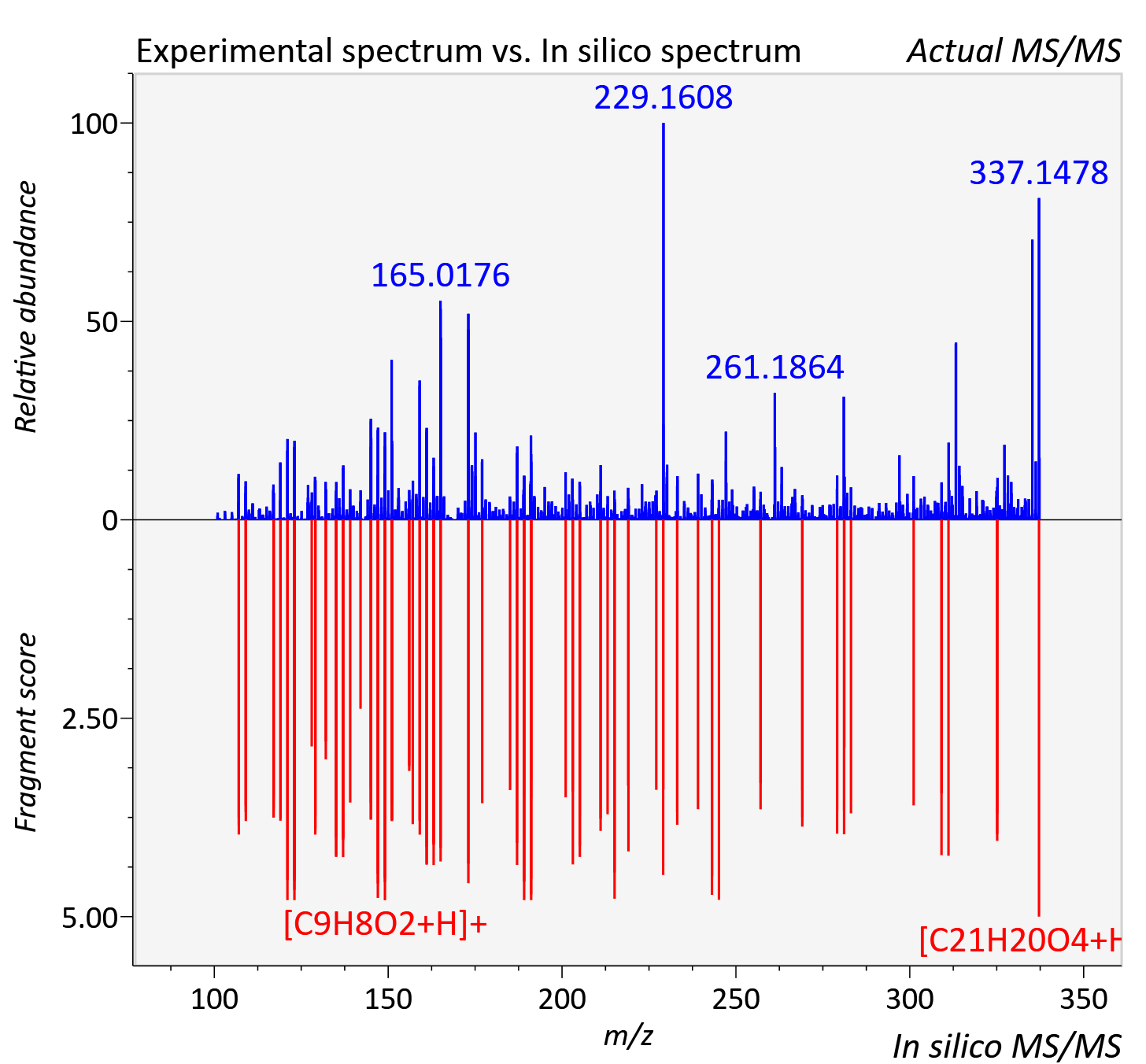
Experimental data
| Retention time | 16.93 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 337.147 | Theoretical mz | 337.143 |
| Error | 9.86 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2247 |
Identifiers and class information
| Inchi key | BOJUBZPEDAAMPG-UXBLZVDNSA-N |
|---|---|
| Smiles | O=C(C=CC1=CC=C2OC(C=CC2=C1OC)(C)C)C3=CC=C(O)C=C3 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Linear 1,3-diarylpropanoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SwissTargetPrediction and SEA |
| P80404 | ABAT | Gamma-amino-N-butyrate transaminase | T59045 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction and SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| P22001 | KCNA3 | Voltage-gated potassium channel subunit Kv1.3 | T76914 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P10636 | MAPT | Microtubule-associated protein tau | T45593 | SEA |
| P19438 | TNFRSF1A | Tumor necrosis factor receptor R1 | T86552 | SEA |
| Q9H4B7 | TUBB1 | Tubulin beta-1 chain | T84397 | SEA |
| P51649 | ALDH5A1 | Succinate semialdehyde dehydrogenase | T13259 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T59045 | DI0060 | Brain cancer | [ICD-11: 2A00] | P80404 | ABAT |
| T59045 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P80404 | ABAT |
| T59045 | DI0135 | Epileptic encephalopathy | [ICD-11: 8A62] | P80404 | ABAT |
| T59045 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P80404 | ABAT |
| T59045 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P80404 | ABAT |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T76914 | DI0198 | Idiopathic inflammatory myopathy | [ICD-11: 4A41] | P22001 | KCNA3 |
| T76914 | DI0351 | Psoriasis | [ICD-11: EA90] | P22001 | KCNA3 |
| T76914 | DI0352 | Psoriatic arthritis | [ICD-11: FA21] | P22001 | KCNA3 |
| T45593 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P10636 | MAPT |
| T45593 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P10636 | MAPT |
| T45593 | DI0265 | Mild neurocognitive disorder | [ICD-11: 6D71] | P10636 | MAPT |
| T86552 | DI0060 | Brain cancer | [ICD-11: 2A00] | P19438 | TNFRSF1A |
| T86552 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P19438 | TNFRSF1A |
| T84397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9H4B7 | TUBB1 |
| T13259 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P51649 | ALDH5A1 |
| T13259 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P51649 | ALDH5A1 |
