Compound details
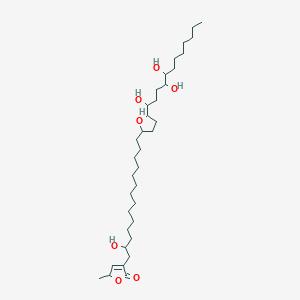
Muricin C
| Compound ID | CDAMM01669 |
|---|---|
| Common name | Muricin C | IUPAC name | 4-[2-hydroxy-14-[5-(1,4,5-trihydroxydodecyl)oxolan-2-yl]tetradecyl]-2-methyl-2H-furan-5-one |
| Molecular formula | C35H64O7 |
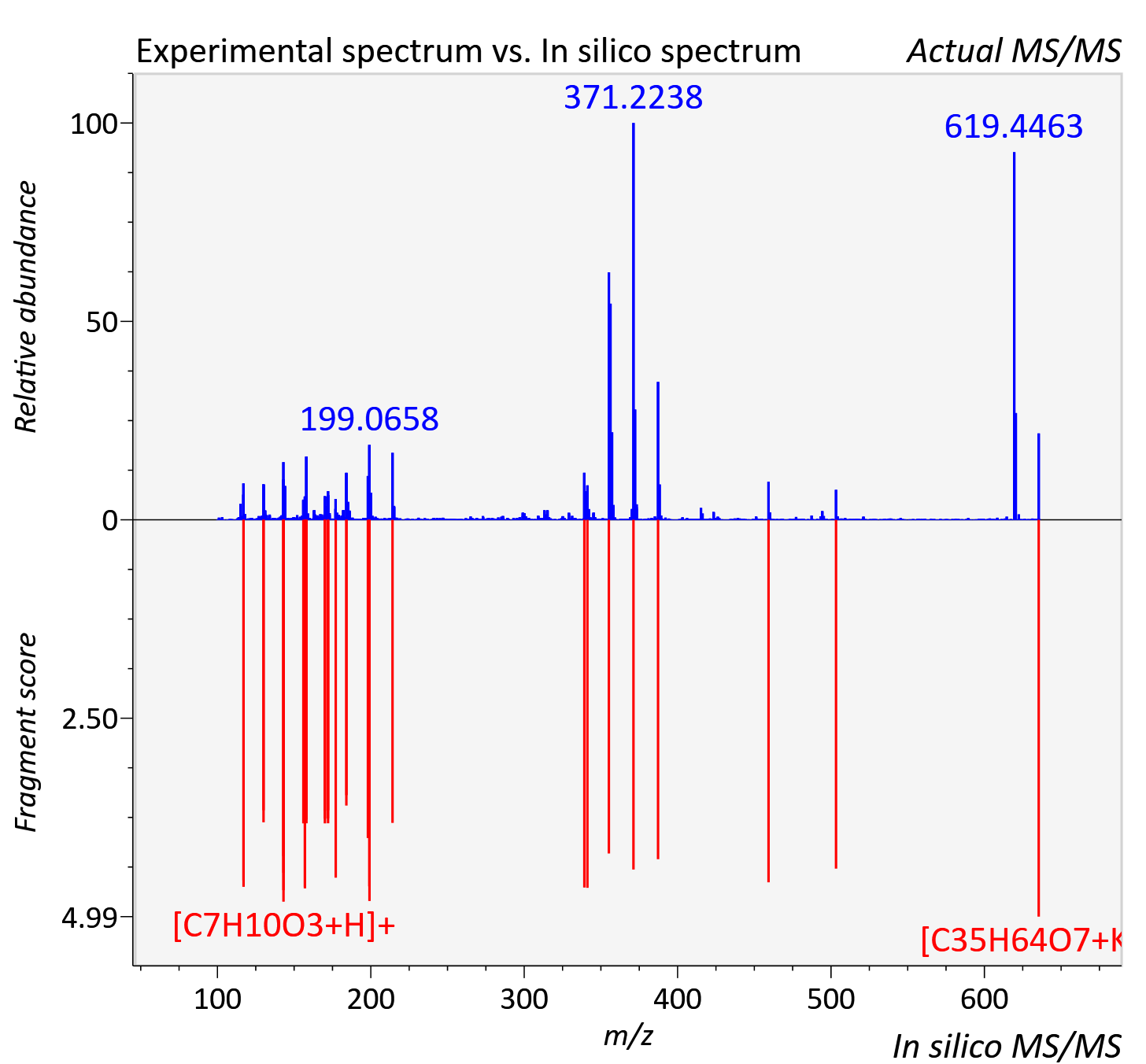
Experimental data
| Retention time | 11.7 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 635.422 | Theoretical mz | 635.428 |
| Error | 9.85 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.3892 |
Identifiers and class information
| Inchi key | PSEVENSSBONHMB-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(C=C1CC(O)CCCCCCCCCCCCC2OC(CC2)C(O)CCC(O)C(O)CCCCCCC)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T62820 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q14416 | GRM2 |
| T62820 | DI0051 | Bipolar disorder | [ICD-11: 6A60] | Q14416 | GRM2 |
| T62820 | DI0116 | Dengue fever | [ICD-11: 1D2Z] | Q14416 | GRM2 |
| T62820 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q14416 | GRM2 |
| T62820 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | Q14416 | GRM2 |
| T62820 | DI0256 | Mental/behavioural/neurodevelopmental disorder | [ICD-11: 6E20-6E8Z] | Q14416 | GRM2 |
| T62820 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | Q14416 | GRM2 |
| T62820 | DI0354 | Psychotic disorder | [ICD-11: 6A20-6A25] | Q14416 | GRM2 |
| T62820 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q14416 | GRM2 |
