Compound details
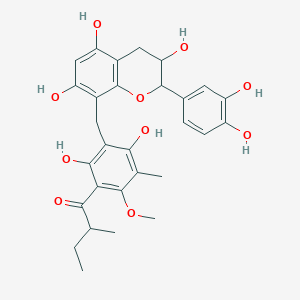
Pilosanol A
| Compound ID | CDAMM01651 |
|---|---|
| Common name | Pilosanol A | IUPAC name | 1-[3-[[2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-3,4-dihydro-2H-chromen-8-yl]methyl]-2,4-dihydroxy-6-methoxy-5-methylphenyl]-2-methylbutan-1-one |
| Molecular formula | C29H32O10 |
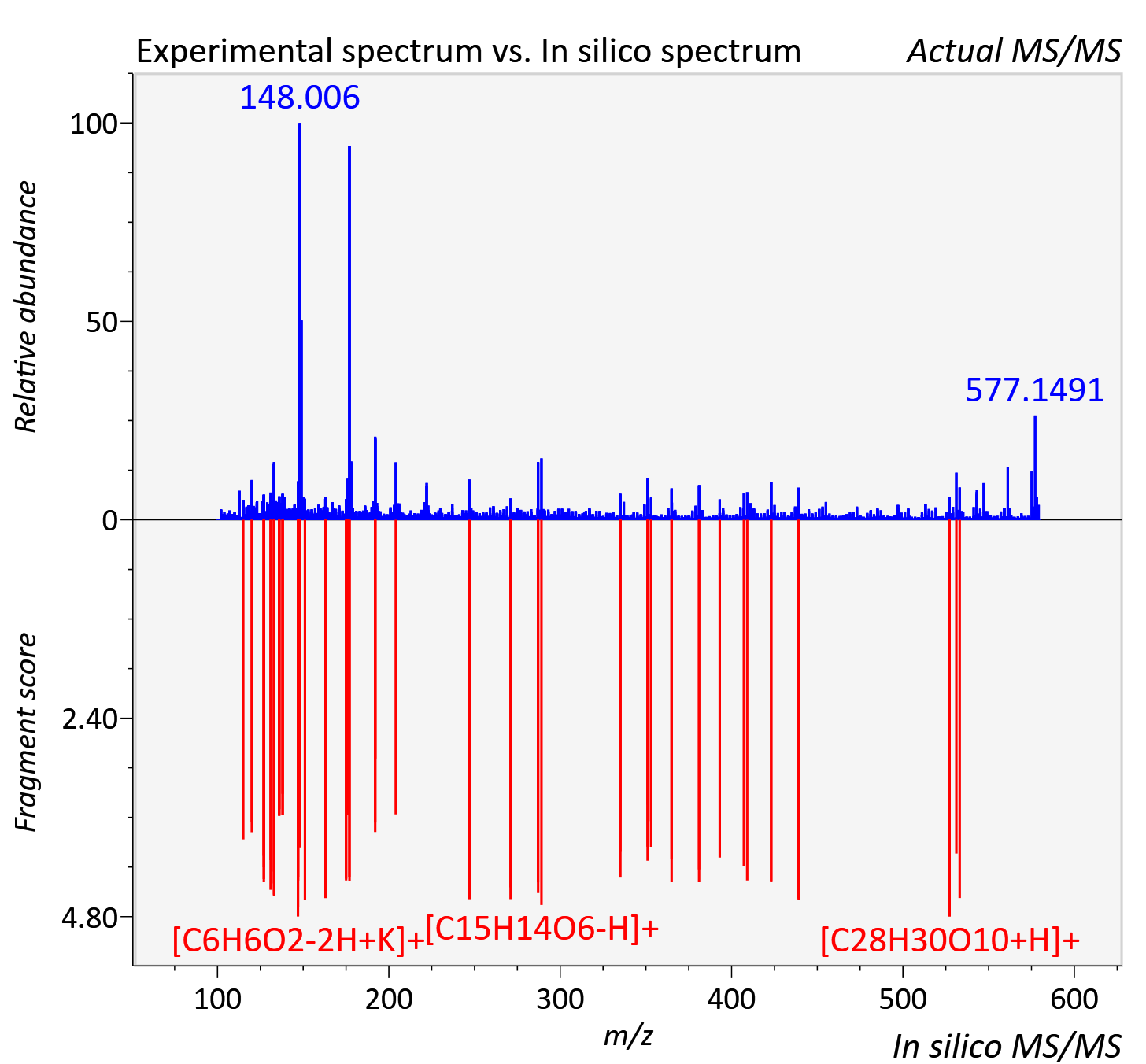
Experimental data
| Retention time | 0.36 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 579.16 | Theoretical mz | 579.163 |
| Error | 5.45 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2749 |
Identifiers and class information
| Inchi key | AXODAJZSINDBPB-QJFKVZSGNA-N |
|---|---|
| Smiles | O=C(C=1C(O)=C(C(O)=C(C1OC)C)CC2=C(O)C=C(O)C3=C2OC(C4=CC=C(O)C(O)=C4)C(O)C3)C(C)CC |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SwissTargetPrediction |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SwissTargetPrediction |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SwissTargetPrediction |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| P08253 | MMP2 | Matrix metalloproteinase 2 | T68251 | SwissTargetPrediction |
| P14780 | MMP9 | Matrix metalloproteinase 9 | T54156 | SwissTargetPrediction |
| Q14534 | SQLE | Squalene monooxygenase (by homology) | T93344 | SwissTargetPrediction |
| P56817 | BACE1 | Beta-secretase 1 | T79031 | SwissTargetPrediction |
| P20151 | KLK2 | Kallikrein 2 | T01908 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SwissTargetPrediction and SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SwissTargetPrediction and SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T68251 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08253 | MMP2 |
| T54156 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P14780 | MMP9 |
| T54156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P14780 | MMP9 |
| T54156 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P14780 | MMP9 |
| T54156 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P14780 | MMP9 |
| T93344 | DI0118 | Dermatophytosis | [ICD-11: 1F28] | Q14534 | SQLE |
| T93344 | DI0385 | Skin fungal infection disorder | [ICD-11: EA60] | Q14534 | SQLE |
| T79031 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P56817 | BACE1 |
| T01908 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P20151 | KLK2 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
