Compound details
Daedaleaside D
| Compound ID | CDAMM01579 |
|---|---|
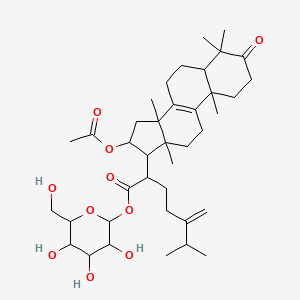
| Common name | Daedaleaside D | IUPAC name | [3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl] 2-(16-acetyloxy-4,4,10,13,14-pentamethyl-3-oxo-1,2,5,6,7,11,12,15,16,17-decahydrocyclopenta[a]phenanthren-17-yl)-6-methyl-5-methylideneheptanoate |
| Molecular formula | C39H60O10 |
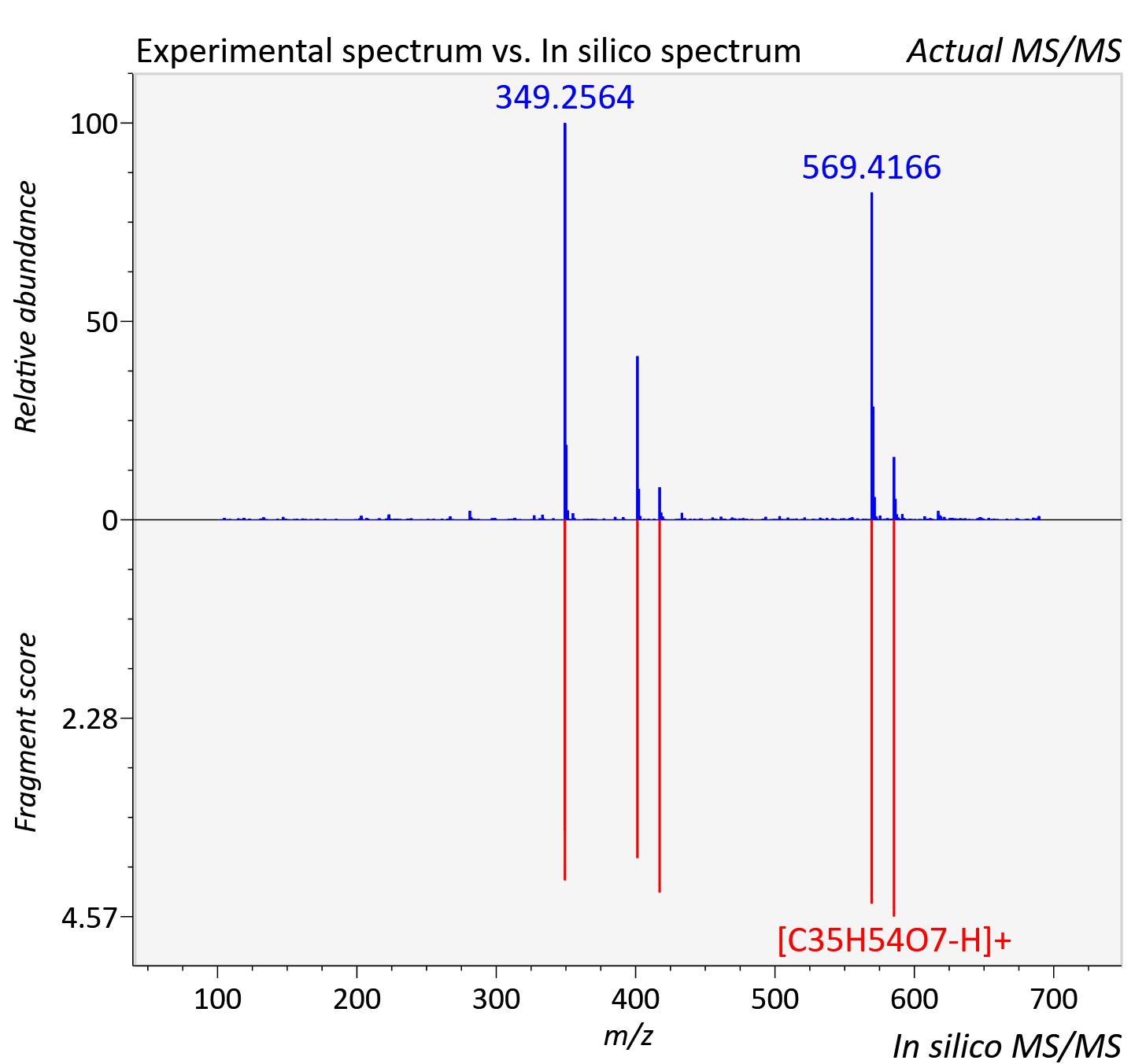
Experimental data
| Retention time | 15.8 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 689.422 | Theoretical mz | 689.426 |
| Error | 6.64 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0416 |
Identifiers and class information
| Inchi key | RSBAXXTZWHYISF-OQWXSBENNA-N |
|---|---|
| Smiles | O=C(OC1CC2(C3=C(CCC2(C)C1C(C(=O)OC4OC(CO)C(O)C(O)C4O)CCC(=C)C(C)C)C5(C)CCC(=O)C(C)(C)C5CC3)C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T94033 | DI0052 | Bleeding disorder | [ICD-11: GA20-GA21] | P00734 | F2 |
| T94033 | DI0091 | Coagulation defect | [ICD-11: 3B10] | P00734 | F2 |
| T94033 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P00734 | F2 |
| T94033 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P00734 | F2 |
| T94033 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P00734 | F2 |
| T94033 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P00734 | F2 |
| T94033 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P00734 | F2 |
| T94033 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P00734 | F2 |
| T16308 | DI0238 | Lung cancer | [ICD-11: 2C25] | Q13526 | PIN1 |
