Compound details
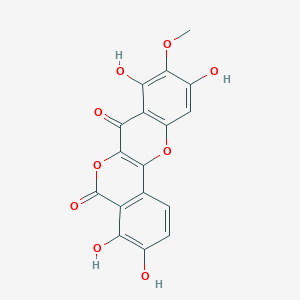
Distemonanthin
| Compound ID | CDAMM01558 |
|---|---|
| Common name | Distemonanthin | IUPAC name | 3,4,8,10-tetrahydroxy-9-methoxyisochromeno[4,3-b]chromene-5,7-dione |
| Molecular formula | C17H10O9 |
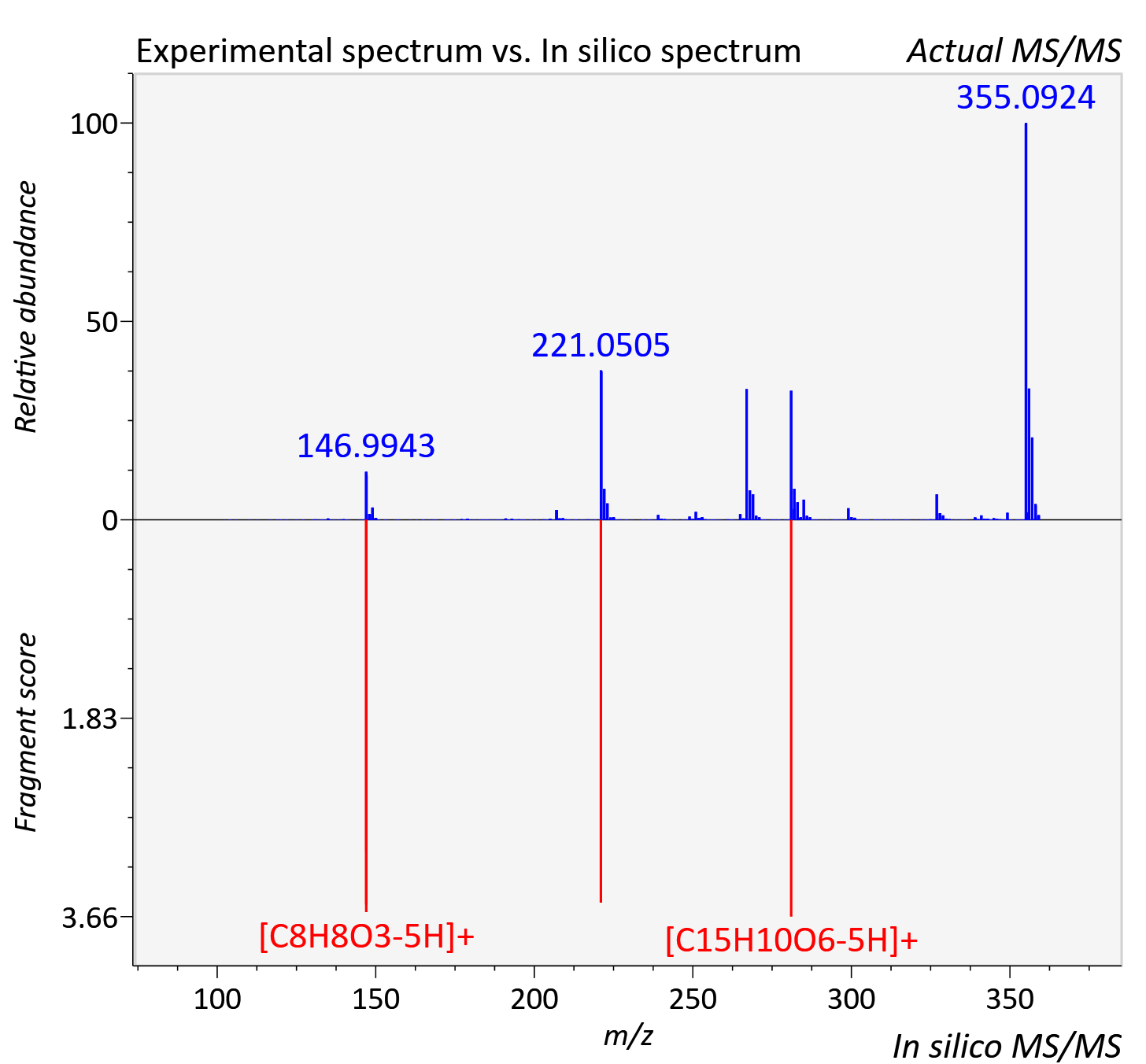
Experimental data
| Retention time | 19.53 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 359.039 | Theoretical mz | 359.039 |
| Error | 0.49 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7417 |
Identifiers and class information
| Inchi key | SLUGCDLLXDGNNT-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C1OC=2C(=O)C3=C(O)C(OC)=C(O)C=C3OC2C=4C=CC(O)=C(O)C14 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SwissTargetPrediction |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SwissTargetPrediction |
| P21397 | MAOA | Monoamine oxidase A | T83875 | SwissTargetPrediction and SEA |
| P00915 | CA1 | Carbonic anhydrase I | T13201 | SwissTargetPrediction |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SwissTargetPrediction and SEA |
| P35354 | PTGS2 | Cyclooxygenase-2 | T66665 | SwissTargetPrediction |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SwissTargetPrediction and SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SwissTargetPrediction and SEA |
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SwissTargetPrediction and SEA |
| Q9HC97 | GPR35 | G-protein coupled receptor 35 | T56143 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P30291 | WEE1 | Serine/threonine-protein kinase WEE1 | T69991 | SwissTargetPrediction |
| P18054 | ALOX12 | Arachidonate 12-lipoxygenase | T35527 | SEA |
| O14746 | TERT | Telomerase reverse transcriptase | T86052 | SEA |
| P00390 | GSR | Glutathione reductase | T30803 | SwissTargetPrediction |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| Q9NPH5 | NOX4 | NADPH oxidase 4 | T29741 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P63316 | TNNC1 | Troponin, cardiac muscle | T12966 | SwissTargetPrediction |
| Q9UNI1 | CELA1 | Elastase 1 | T22839 | SEA |
| Q9HC98 | NEK6 | Serine/threonine-protein kinase NEK6 | T78992 | SEA |
| O60285 | NUAK1 | NUAK family SNF1-like kinase 1 | T21653 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SwissTargetPrediction and SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q9NZ08 | ERAP1 | Endoplasmic reticulum aminopeptidase 1 | T72849 | SEA |
| P05231 | IL6 | Interleukin-6 | T32578 | SEA |
| O35273 | SMAD3 | Mothers against decapentaplegic homolog 3 | T35445 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
| P16220 | CREB1 | Cyclic AMP-responsive element-binding protein | T92098 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
| Q9UK17 | KCND3 | Voltage-gated potassium channel Kv4.3 | T74500 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T83875 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P21397 | MAOA |
| T83875 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P21397 | MAOA |
| T13201 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00915 | CA1 |
| T13201 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00915 | CA1 |
| T66665 | DI0087 | Chronic pain | [ICD-11: MG30] | P35354 | PTGS2 |
| T66665 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P35354 | PTGS2 |
| T66665 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P35354 | PTGS2 |
| T66665 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P35354 | PTGS2 |
| T66665 | DI0227 | Knee osteoarthritis | [ICD-11: FA01] | P35354 | PTGS2 |
| T66665 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35354 | PTGS2 |
| T66665 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35354 | PTGS2 |
| T66665 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P35354 | PTGS2 |
| T66665 | DI0416 | Tuberculosis | [ICD-11: 1B10-1B12] | P35354 | PTGS2 |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T69991 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P30291 | WEE1 |
| T69991 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P30291 | WEE1 |
| T69991 | DI0429 | Uterine ligament/parametrium/uterine adnexa neoplasm | [ICD-11: 2C72] | P30291 | WEE1 |
| T86052 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | O14746 | TERT |
| T86052 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | O14746 | TERT |
| T86052 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | O14746 | TERT |
| T30803 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P00390 | GSR |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T29741 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9NPH5 | NOX4 |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T12966 | DI0006 | Acquired cutaneous blood vessel malformation | [ICD-11: EF20] | P63316 | TNNC1 |
| T12966 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P63316 | TNNC1 |
| T22839 | DI0024 | Alpha-1-antitrypsin deficiency | [ICD-11: 5C5A] | Q9UNI1 | CELA1 |
| T22839 | DI0065 | Bronchitis | [ICD-11: CA20] | Q9UNI1 | CELA1 |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T32578 | DI0028 | Anemia | [ICD-11: 3A00-3A9Z] | P05231 | IL6 |
| T35445 | DI0226 | Kidney fibrosis | [ICD-11: GC01] | O35273 | SMAD3 |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
| T74500 | DI0004 | Acidosis | [ICD-11: 5C73] | Q9UK17 | KCND3 |
