Compound details
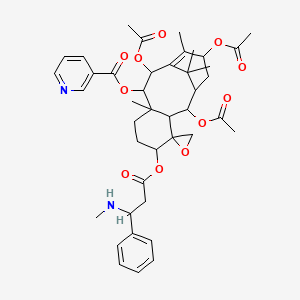
N-Demethylnicaustrine
| Compound ID | CDAMM01521 |
|---|---|
| Common name | N-Demethylnicaustrine | IUPAC name | [2\',10\',13\'-triacetyloxy-8\',12\',15\',15\'-tetramethyl-5\'-[3-(methylamino)-3-phenylpropanoyl]oxyspiro[oxirane-2,4\'-tricyclo[9.3.1.03,8]pentadec-11-ene]-9\'-yl] pyridine-3-carboxylate |
| Molecular formula | C42H52N2O11 |
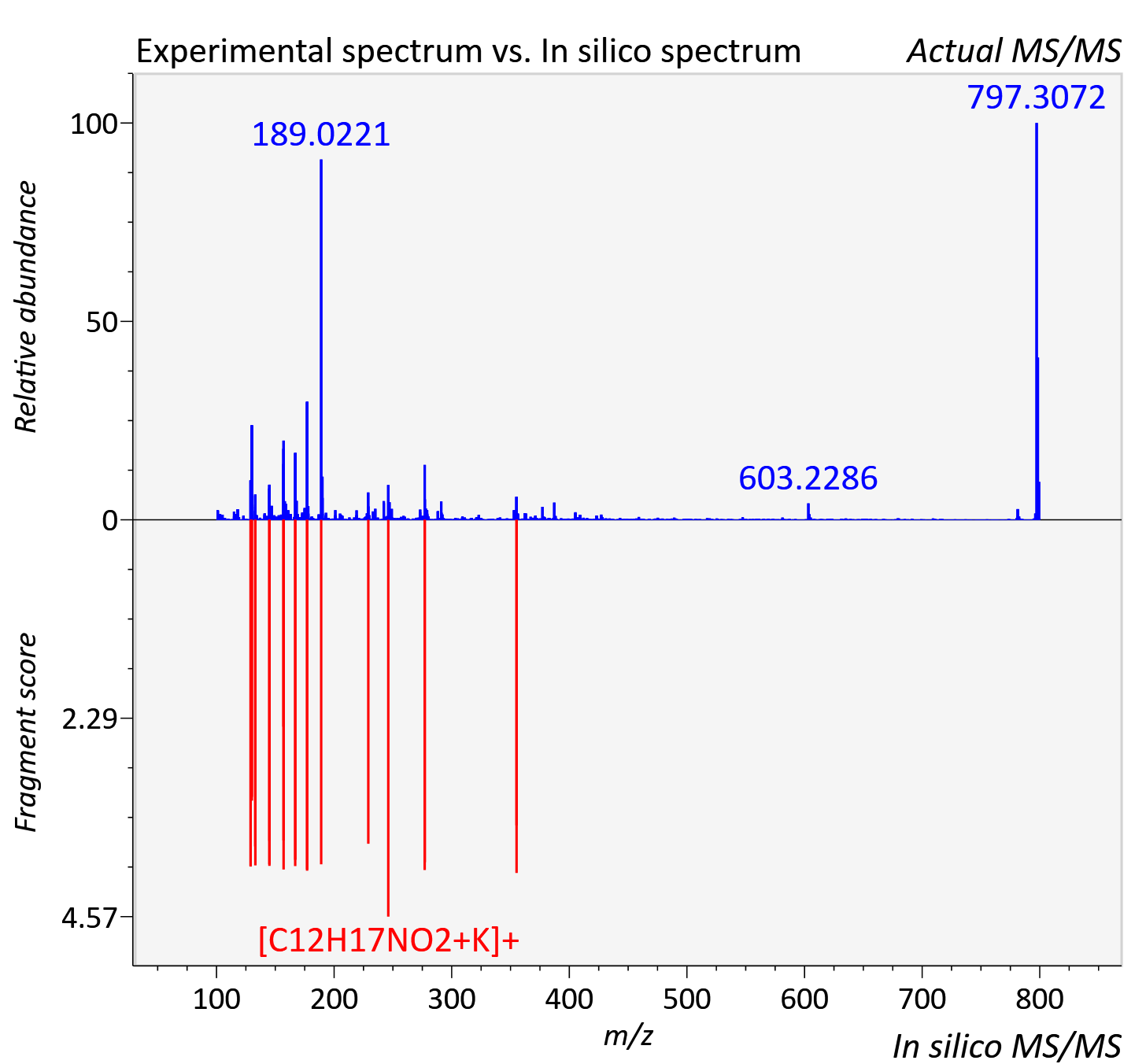
Experimental data
| Retention time | 7.8 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 799.315 | Theoretical mz | 799.32 |
| Error | 6.95 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.1629 |
Identifiers and class information
| Inchi key | MYTZQCJTTCFLHG-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(OC1C(OC(=O)C)C2=C(C)C(OC(=O)C)CC(C(OC(=O)C)C3C4(OC4)C(OC(=O)CC(NC)C=5C=CC=CC5)CCC13C)C2(C)C)C=6C=NC=CC6 |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| O75908 | SOAT2 | Acyl coenzyme A:cholesterol acyltransferase 2 | T14463 | SEA |
| O75908 | SOAT2 | Acyl coenzyme A:cholesterol acyltransferase 2 | T14463 | SEA |
| P40306 | PSMB10 | 26S proteosome | T09022 | SEA |
| P25786 | PSMA1 | Core proteasome 20S complex | T61997 | SEA |
| P28072 | PSMB6 | Proteasome beta-1 | T54973 | SEA |
| Q99436 | PSMB7 | Proteasome beta-2 | T55986 | SEA |
