Compound details
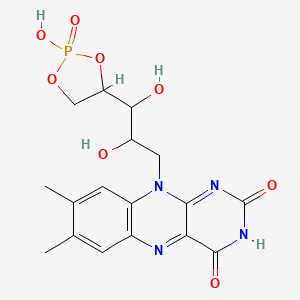
Riboflavin cyclic-4\',5\'-phosphate
| Compound ID | CDAMM01505 |
|---|---|
| Common name | Riboflavin cyclic-4\',5\'-phosphate | IUPAC name | 10-[2,3-dihydroxy-3-(2-hydroxy-2-oxo-1,3,2lambda5-dioxaphospholan-4-yl)propyl]-7,8-dimethylbenzo[g]pteridine-2,4-dione |
| Molecular formula | C17H19N4O8P |
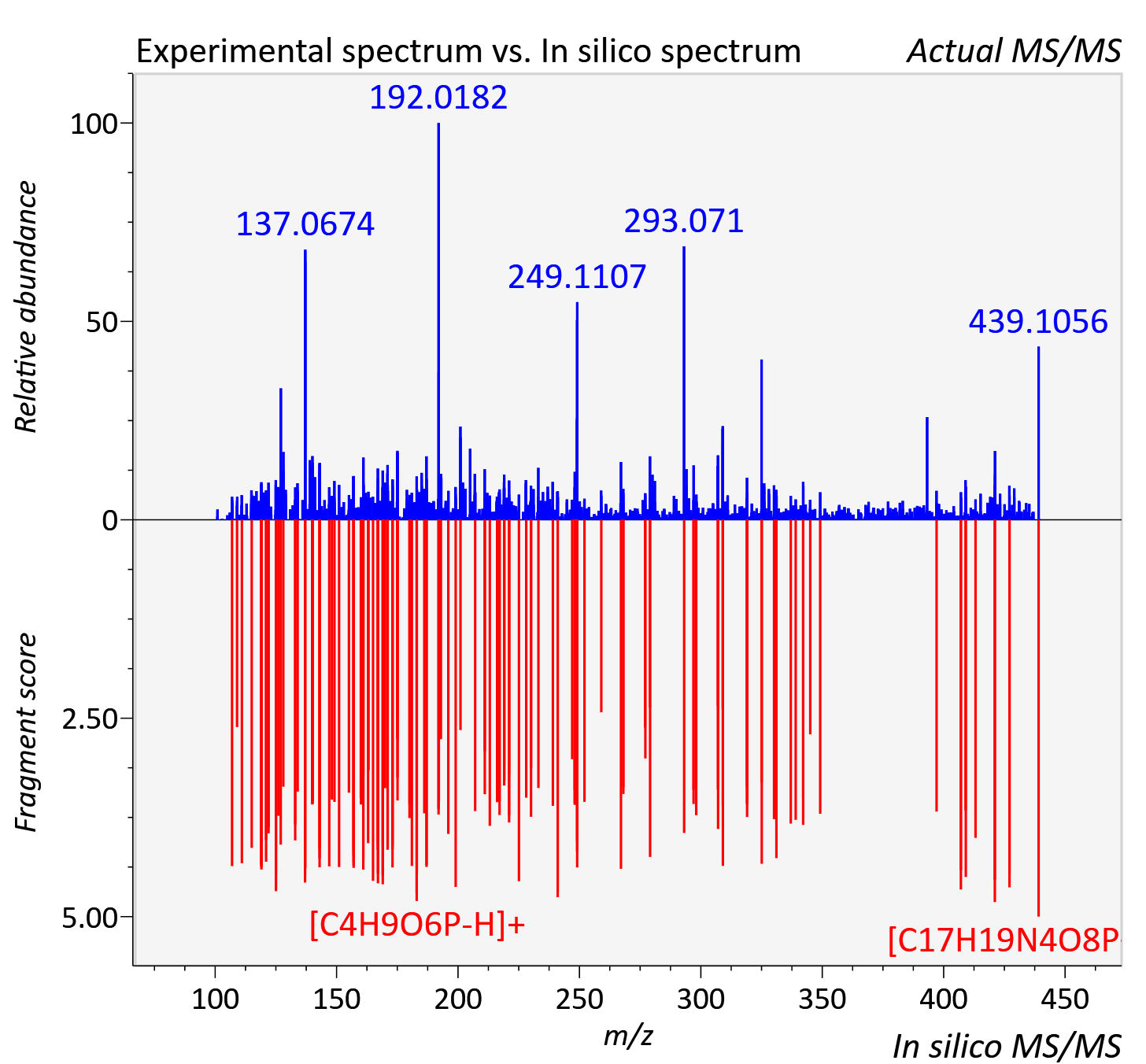
Experimental data
| Retention time | 2.89 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 439.102 | Theoretical mz | 439.101 |
| Error | 1.65 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.6371 |
Identifiers and class information
| Inchi key | CVZKYDYRJQYYDJ-QTZZOOGMNA-N |
|---|---|
| Smiles | O=C1N=C(O)C2=NC3=CC(=C(C=C3N(C2=N1)CC(O)C(O)C4OP(=O)(O)OC4)C)C |
| Superclass | Organoheterocyclic compounds |
| Class | Pteridines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P30542 | ADORA1 | Adenosine A1 receptor | T92072 | SwissTargetPrediction |
| P08254 | MMP3 | Matrix metalloproteinase 3 | T86702 | SwissTargetPrediction |
| P14780 | MMP9 | Matrix metalloproteinase 9 | T54156 | SwissTargetPrediction |
| P03956 | MMP1 | Matrix metalloproteinase 1 | T52450 | SwissTargetPrediction |
| P07195 | LDHB | L-lactate dehydrogenase B chain | T88057 | SwissTargetPrediction |
| P78536 | ADAM17 | ADAM17 | T82393 | SwissTargetPrediction |
| P25101 | EDNRA | Endothelin receptor ET-A | T23499 | SwissTargetPrediction |
| P20839 | IMPDH1 | Inosine-5'-monophosphate dehydrogenase 1 | T40111 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T92072 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P30542 | ADORA1 |
| T86702 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P08254 | MMP3 |
| T86702 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P08254 | MMP3 |
| T86702 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P08254 | MMP3 |
| T86702 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P08254 | MMP3 |
| T86702 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P08254 | MMP3 |
| T86702 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P08254 | MMP3 |
| T86702 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P08254 | MMP3 |
| T54156 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P14780 | MMP9 |
| T54156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P14780 | MMP9 |
| T54156 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P14780 | MMP9 |
| T54156 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P14780 | MMP9 |
| T52450 | DI0238 | Lung cancer | [ICD-11: 2C25] | P03956 | MMP1 |
| T82393 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P78536 | ADAM17 |
| T82393 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P78536 | ADAM17 |
| T23499 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P25101 | EDNRA |
| T23499 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P25101 | EDNRA |
| T23499 | DI0425 | Urinary system clinical symptom | [ICD-11: MF8Y] | P25101 | EDNRA |
| T40111 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P20839 | IMPDH1 |
| T40111 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P20839 | IMPDH1 |
| T40111 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | P20839 | IMPDH1 |
| T40111 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P20839 | IMPDH1 |
| T40111 | DI0413 | Transplant rejection | [ICD-11: NE84] | P20839 | IMPDH1 |
