Compound details
Fukiic acid
| Compound ID | CDAMM01496 |
|---|---|
| Common name | Fukiic acid | IUPAC name | 2-[(3,4-dihydroxyphenyl)methyl]-2,3-dihydroxybutanedioic acid |
| Molecular formula | C11H12O8 |
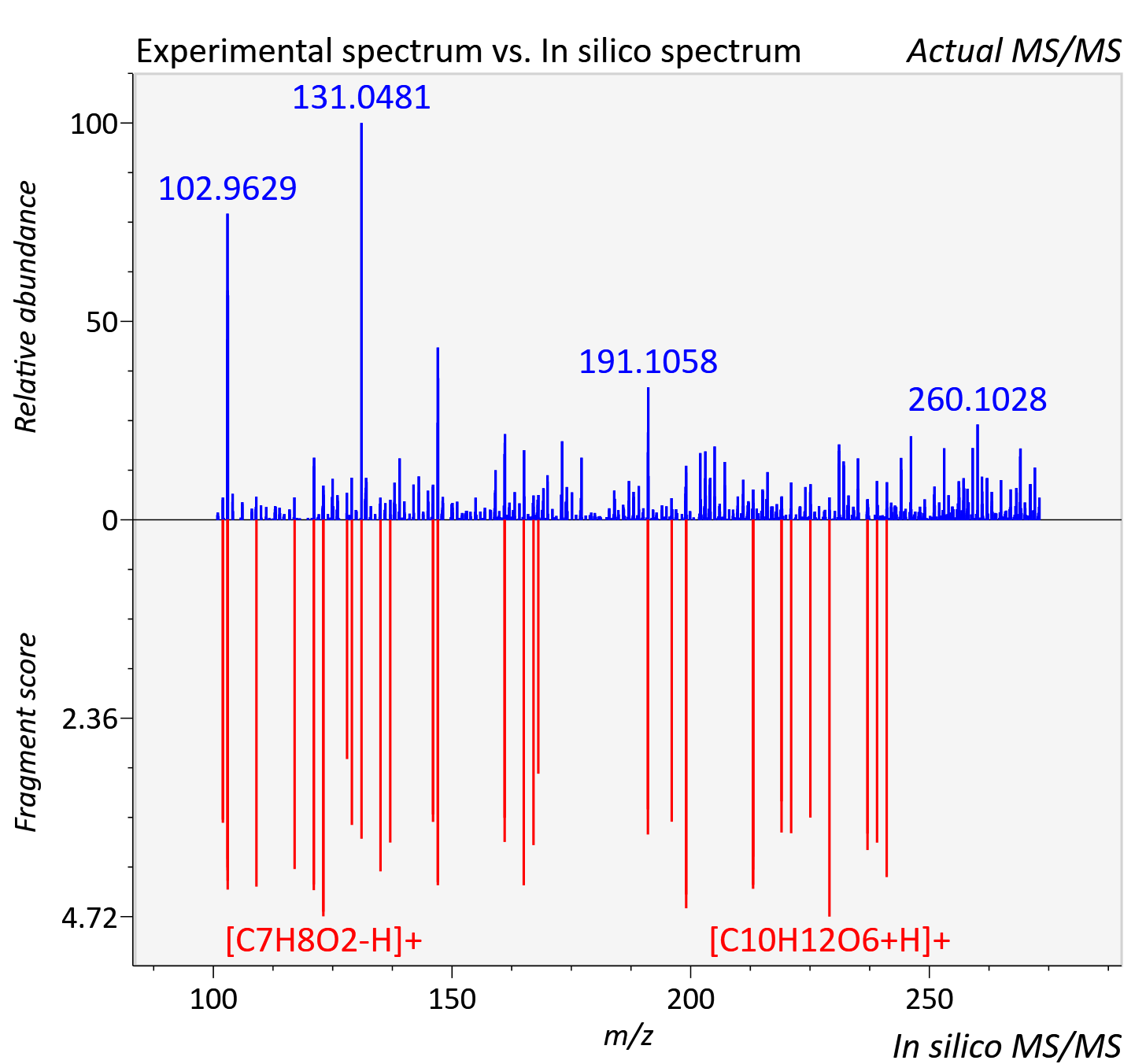
Experimental data
| Retention time | 11.42 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 273.061 | Theoretical mz | 273.06 |
| Error | 3.38 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.2917 |
Identifiers and class information
| Inchi key | PHFSBARLASYIFM-LDYMZIIASA-N |
|---|---|
| Smiles | O=C(O)C(O)C(O)(C(=O)O)CC1=CC=C(O)C(O)=C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Phenylpropanoic acids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P06241 | FYN | Tyrosine-protein kinase FYN | T17980 | SEA |
| P02766 | TTR | Transthyretin | T86462 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P08908 | HTR1A | Serotonin 1a (5-HT1a) receptor | T78709 | SwissTargetPrediction |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| Q96IY4 | CPB2 | Carboxypeptidase B2 isoform A | T98022 | SwissTargetPrediction |
| P07101 | TH | Tyrosine 3-hydroxylase | T62390 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SwissTargetPrediction |
| Q01650 | SLC7A5 | L-type amino acid transporter 1 (by homology) | T48330 | SEA |
| P16050 | ALOX15 | Arachidonate 15-lipoxygenase | T16042 | SEA |
| Q02750 | MAP2K1 | Dual specificity mitogen-activated protein kinase kinase 1 | T99057 | SwissTargetPrediction |
| P15085 | CPA1 | Carboxypeptidase A1 | T33966 | SwissTargetPrediction |
| Q86YT5 | SLC13A5 | Solute carrier family 13 member 5 | T81547 | SwissTargetPrediction and SEA |
| P17936 | IGFBP3 | Insulin-like growth factor binding protein 3 | T33455 | SEA |
| P53396 | ACLY | ATP-citrate synthase | T52189 | SEA |
| P24593 | IGFBP5 | Insulin-like growth factor binding protein 5 | T17008 | SEA |
| P09172 | DBH | Dopamine beta hydroxylase | T74937 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T17980 | DI0286 | Myeloproliferative neoplasm | [ICD-11: 2A20] | P06241 | FYN |
| T86462 | DI0026 | Amyloidosis | [ICD-11: 5D00] | P02766 | TTR |
| T78709 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | P08908 | HTR1A |
| T78709 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P08908 | HTR1A |
| T78709 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P08908 | HTR1A |
| T78709 | DI0264 | Migraine | [ICD-11: 8A80] | P08908 | HTR1A |
| T78709 | DI0270 | Mood disorder | [ICD-11: 6A60-6E23] | P08908 | HTR1A |
| T98022 | DI0073 | Cerebral ischaemia | [ICD-11: 8B1Z] | Q96IY4 | CPB2 |
| T98022 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q96IY4 | CPB2 |
| T98022 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | Q96IY4 | CPB2 |
| T62390 | DI0019 | Adrenomedullary hyperfunction | [ICD-11: 5A75] | P07101 | TH |
| T62390 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P07101 | TH |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T99057 | DI0262 | Metastatic tumour | [ICD-11: 2D50-2E2Z] | Q02750 | MAP2K1 |
| T99057 | DI0336 | Phakomatoses/hamartoneoplastic syndrome | [ICD-11: LD2D] | Q02750 | MAP2K1 |
| T33455 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P17936 | IGFBP3 |
| T52189 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P53396 | ACLY |
| T74937 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P09172 | DBH |
| T74937 | DI0341 | Post-traumatic stress disorder | [ICD-11: 6B40] | P09172 | DBH |
