Compound details
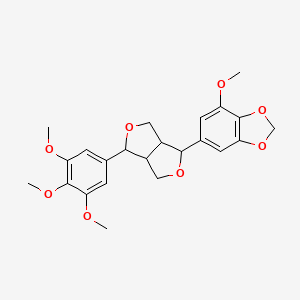
Sesartemin
| Compound ID | CDAMM01445 |
|---|---|
| Common name | Sesartemin | IUPAC name | 4-methoxy-6-[6-(3,4,5-trimethoxyphenyl)-1,3,3a,4,6,6a-hexahydrofuro[3,4-c]furan-3-yl]-1,3-benzodioxole |
| Molecular formula | C23H26O8 |
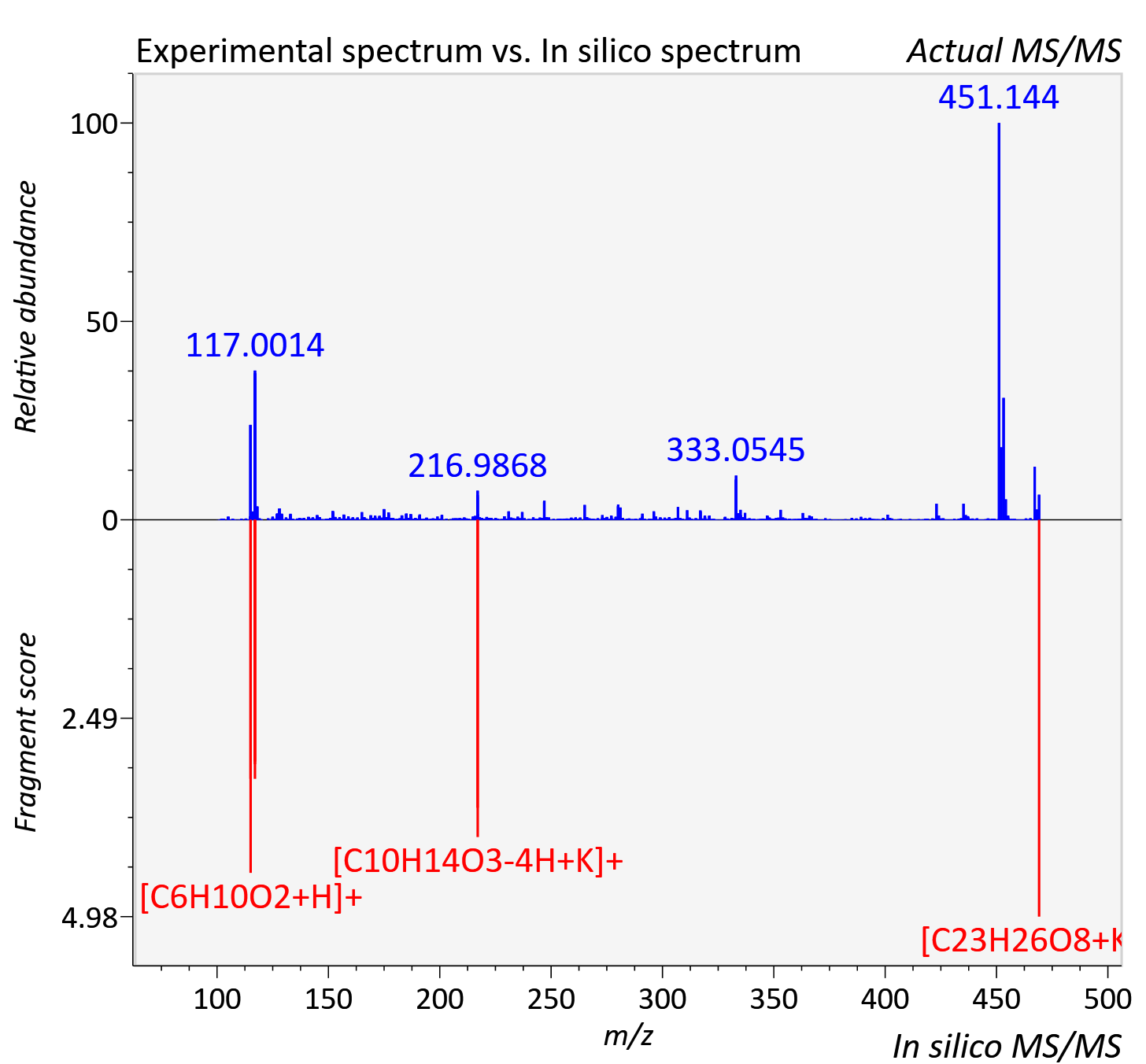
Experimental data
| Retention time | 3.74 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 469.129 | Theoretical mz | 469.126 |
| Error | 5.42 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7293 |
Identifiers and class information
| Inchi key | DHWUVPPRBIJJKS-UHFFFAOYNA-N |
|---|---|
| Smiles | O(C=1C=C(C=C(OC)C1OC)C2OCC3C(OCC23)C4=CC(OC)=C5OCOC5=C4)C |
| Superclass | Lignans, neolignans and related compounds |
| Class | Furanoid lignans |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q07820 | MCL1 | Induced myeloid leukemia cell differentiation protein Mcl-1 | T14912 | SwissTargetPrediction |
| P53779 | MAPK10 | c-Jun N-terminal kinase 3 | T85421 | SwissTargetPrediction |
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SwissTargetPrediction |
| O00206 | TLR4 | Toll-like receptor 4 (by homology) | T81443 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P25105 | PTAFR | Platelet activating factor receptor | T87023 | SwissTargetPrediction and SEA |
| P25101 | EDNRA | Endothelin receptor ET-A | T23499 | SEA |
| P24530 | EDNRB | Endothelin receptor ET-B | T92828 | SEA |
| P00746 | CFD | Complement factor D | T66383 | SwissTargetPrediction |
| P51843 | NR0B1 | Orphan nuclear receptor DAX-1 | T23191 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| Q9P0J0 | NDUFA13 | Mitochondrial complex I | T82391 | SEA |
| Q9Y6M9 | NDUFB9 | HUMAN NADH:ubiquinone oxidoreductase subunit B9 | T01465 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| P03897 | MT-ND3 | NADH dehydrogenase | T77195 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T14912 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q07820 | MCL1 |
| T14912 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | Q07820 | MCL1 |
| T14912 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q07820 | MCL1 |
| T14912 | DI0252 | Melanoma | [ICD-11: 2C30] | Q07820 | MCL1 |
| T14912 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | Q07820 | MCL1 |
| T14912 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q07820 | MCL1 |
| T85421 | DI0125 | Dissociative neurological symptom disorder | [ICD-11: 6B60] | P53779 | MAPK10 |
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T81443 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | O00206 | TLR4 |
| T81443 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O00206 | TLR4 |
| T81443 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | O00206 | TLR4 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T87023 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P25105 | PTAFR |
| T23499 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P25101 | EDNRA |
| T23499 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P25101 | EDNRA |
| T23499 | DI0425 | Urinary system clinical symptom | [ICD-11: MF8Y] | P25101 | EDNRA |
| T92828 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P24530 | EDNRB |
| T92828 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P24530 | EDNRB |
| T66383 | DI0170 | Haemolytic anemia | [ICD-11: 3A20-3A2Z] | P00746 | CFD |
| T66383 | DI0365 | Retinopathy | [ICD-11: 9B71] | P00746 | CFD |
| T82391 | DI0069 | Cardiomyopathy | [ICD-11: BC43] | Q9P0J0 | NDUFA13 |
| T77195 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P03897 | MT-ND3 |
