Compound details
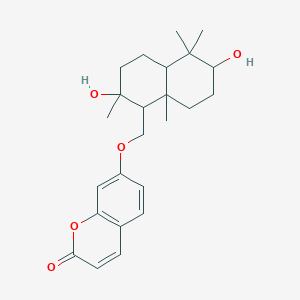
Nevskin
| Compound ID | CDAMM01421 |
|---|---|
| Common name | Nevskin | IUPAC name | 7-[(2,6-dihydroxy-2,5,5,8a-tetramethyl-3,4,4a,6,7,8-hexahydro-1H-naphthalen-1-yl)methoxy]chromen-2-one |
| Molecular formula | C24H32O5 |
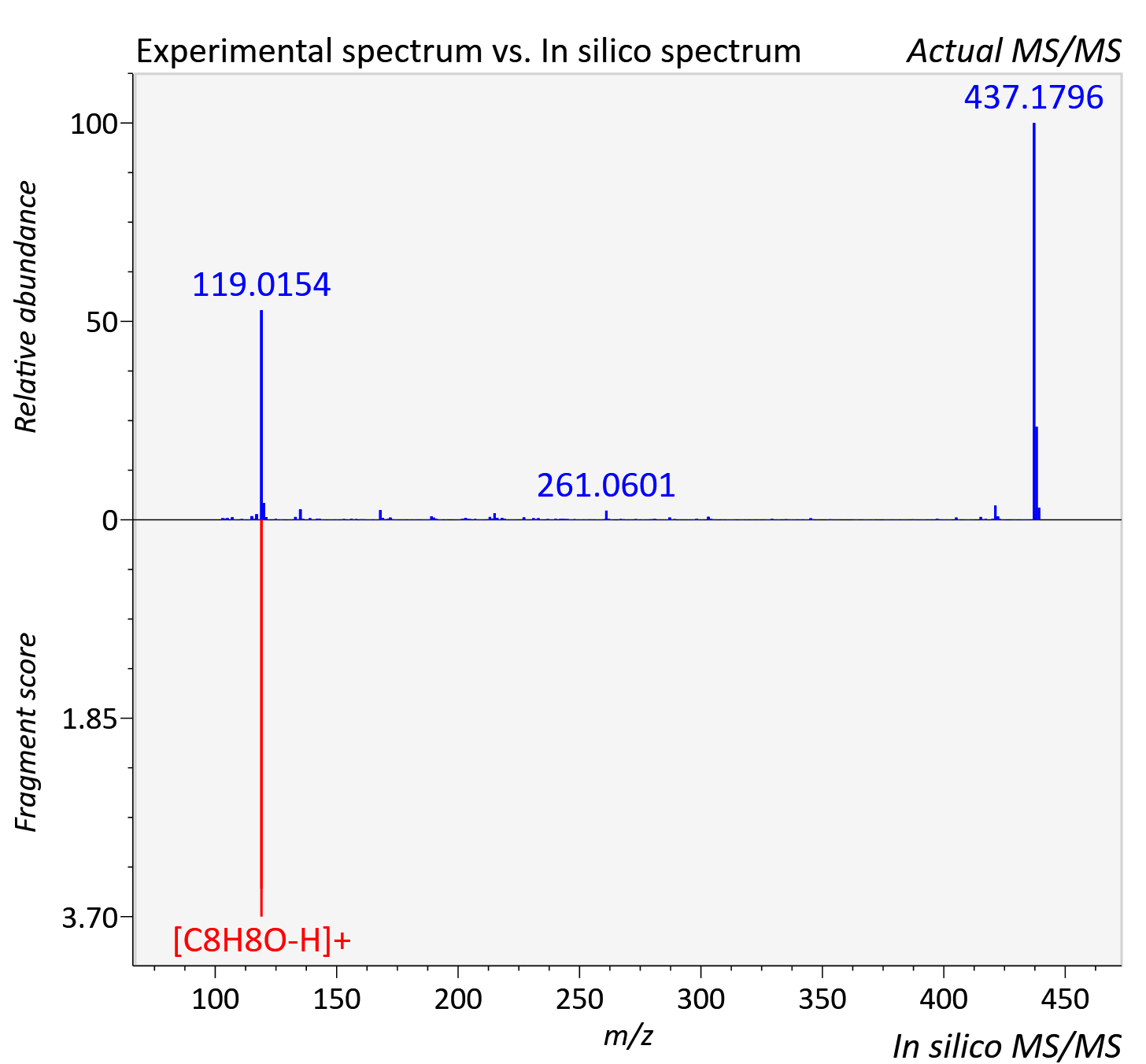
Experimental data
| Retention time | 8.31 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 439.188 | Theoretical mz | 439.188 |
| Error | 1.5 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7524 |
Identifiers and class information
| Inchi key | WNANPKYNOALKIV-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC=2C=C(OCC3C(O)(C)CCC4C(C)(C)C(O)CCC34C)C=CC2C=C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Coumarins and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SwissTargetPrediction and SEA |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SwissTargetPrediction and SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SwissTargetPrediction and SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SEA |
| P17706 | PTPN2 | T-cell protein-tyrosine phosphatase | T49156 | SEA |
| P06276 | BCHE | Butyrylcholinesterase | T99799 | SEA |
| P01584 | IL1B | Interleukin-1 beta | T42000 | SEA |
| Q15119 | PDK2 | Pyruvate dehydrogenase kinase isoform 2 | T17967 | SEA |
| Q9NZ08 | ERAP1 | Endoplasmic reticulum aminopeptidase 1 | T72849 | SEA |
| O15427 | SLC16A3 | Monocarboxylate transporter 4 | T31788 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T49156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P17706 | PTPN2 |
| T99799 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P06276 | BCHE |
| T99799 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P06276 | BCHE |
| T42000 | DI0267 | Mineral excesses | [ICD-11: 5B91] | P01584 | IL1B |
| T42000 | DI0269 | Monogenic autoinflammatory syndrome | [ICD-11: 4A60] | P01584 | IL1B |
| T42000 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P01584 | IL1B |
| T42000 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P01584 | IL1B |
| T17967 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | Q15119 | PDK2 |
