Compound details
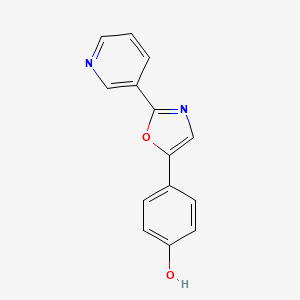
Halfordinol
| Compound ID | CDAMM01419 |
|---|---|
| Common name | Halfordinol | IUPAC name | 4-(2-pyridin-3-yl-1,3-oxazol-5-yl)phenol |
| Molecular formula | C14H10N2O2 |
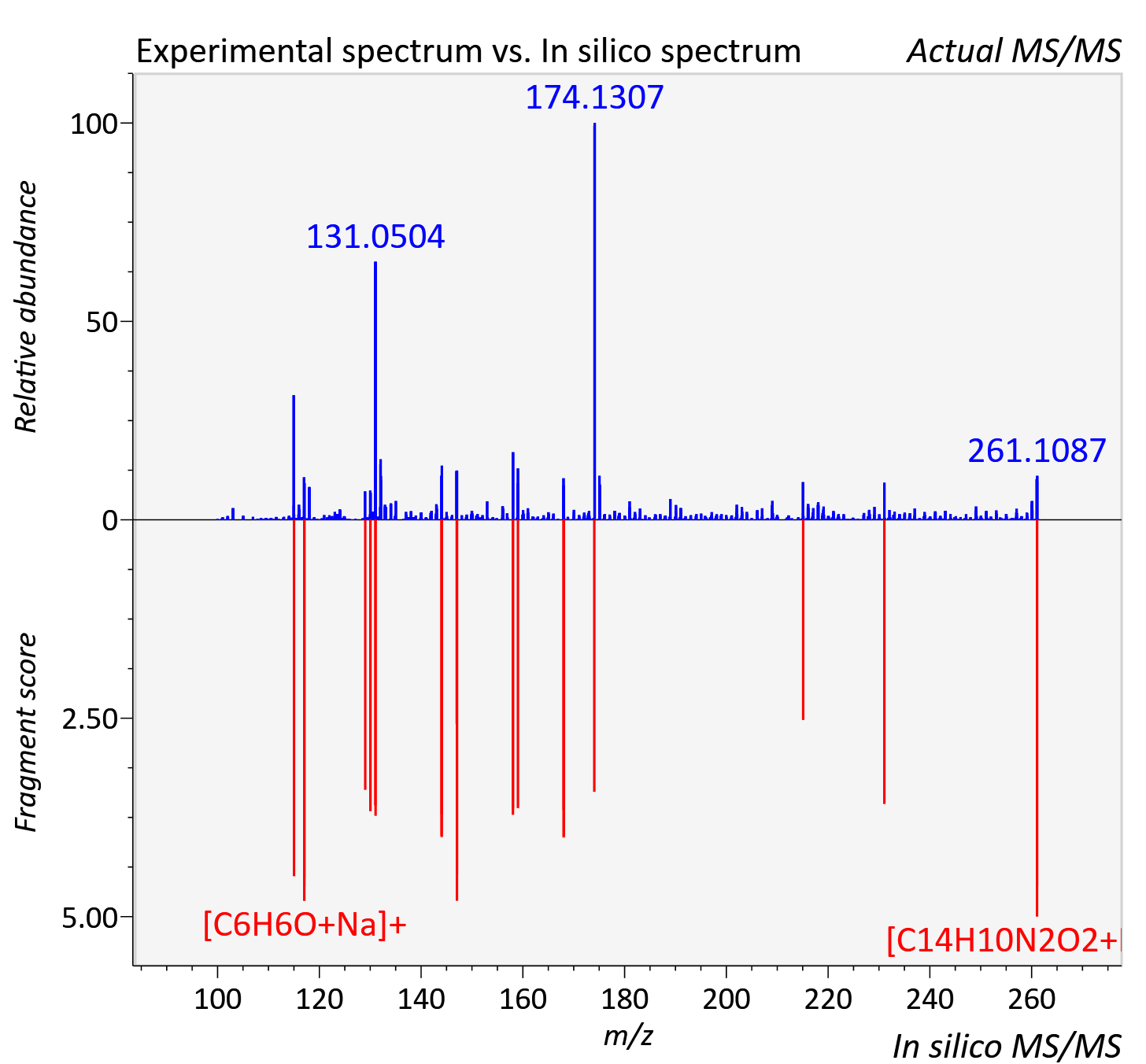
Experimental data
| Retention time | 8.47 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 261.06 | Theoretical mz | 261.063 |
| Error | 12.33 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2401 |
Identifiers and class information
| Inchi key | FUXBKWOAAPDDGE-UHFFFAOYSA-N |
|---|---|
| Smiles | OC1=CC=C(C=C1)C=2OC(=NC2)C3=CN=CC=C3 |
| Superclass | Organoheterocyclic compounds |
| Class | Azoles |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P11509 | CYP2A6 | Cytochrome P450 2A6 | T06455 | SEA |
| P15538 | CYP11B1 | Cytochrome P450 11B1 | T84621 | SEA |
| P19099 | CYP11B2 | Cytochrome P450 11B2 | T59056 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SEA |
| P55072 | VCP | Transitional endoplasmic reticulum ATPase | T12646 | SEA |
| P05093 | CYP17A1 | Cytochrome P450 17A1 | T89041 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SEA |
| O60760 | HPGDS | Hematopoietic prostaglandin D synthase | T19433 | SEA |
| P42224 | STAT1 | Signal transducer and activator of transcription 1-alpha/beta | T64205 | SEA |
| P40337 | VHL | Von Hippel-Lindau disease tumor suppressor/Elongin B/Elongin C | T80423 | SEA |
| Q9Y337 | KLK5 | Kallikrein 5 | T11808 | SEA |
| P30532 | CHRNA5 | Nicotinic acetylcholine receptor alpha4/beta2/alpha5 | T00299 | SEA |
| Q9P0G3 | KLK14 | Kallikrein-14 | T39195 | SEA |
| P20813 | CYP2B6 | Cytochrome P450 2B6 | T11793 | SEA |
| Q96HK3 | CALM | Calmodulin | T39610 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T06455 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P11509 | CYP2A6 |
| T06455 | DI0351 | Psoriasis | [ICD-11: EA90] | P11509 | CYP2A6 |
| T84621 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P15538 | CYP11B1 |
| T84621 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P15538 | CYP11B1 |
| T59056 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P19099 | CYP11B2 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T12646 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P55072 | VCP |
| T12646 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | P55072 | VCP |
| T12646 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P55072 | VCP |
| T12646 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P55072 | VCP |
| T89041 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P05093 | CYP17A1 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T19433 | DI0148 | Flatworm infection | [ICD-11: 1F70-1F86] | O60760 | HPGDS |
| T64205 | DI0383 | Skin and skin-structure infection | [ICD-11: 1F28-1G0Z] | P42224 | STAT1 |
| T39610 | DI0068 | Cardiac arrhythmia | [ICD-11: BC9Z] | Q96HK3 | CALM |
| T39610 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q96HK3 | CALM |
| T39610 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q96HK3 | CALM |
