Compound details
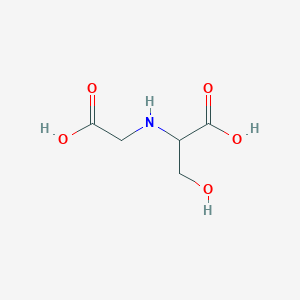
L-N-Carboxymethylserine
| Compound ID | CDAMM01417 |
|---|---|
| Common name | L-N-Carboxymethylserine | IUPAC name | 2-(carboxymethylamino)-3-hydroxypropanoic acid |
| Molecular formula | C5H9NO5 |
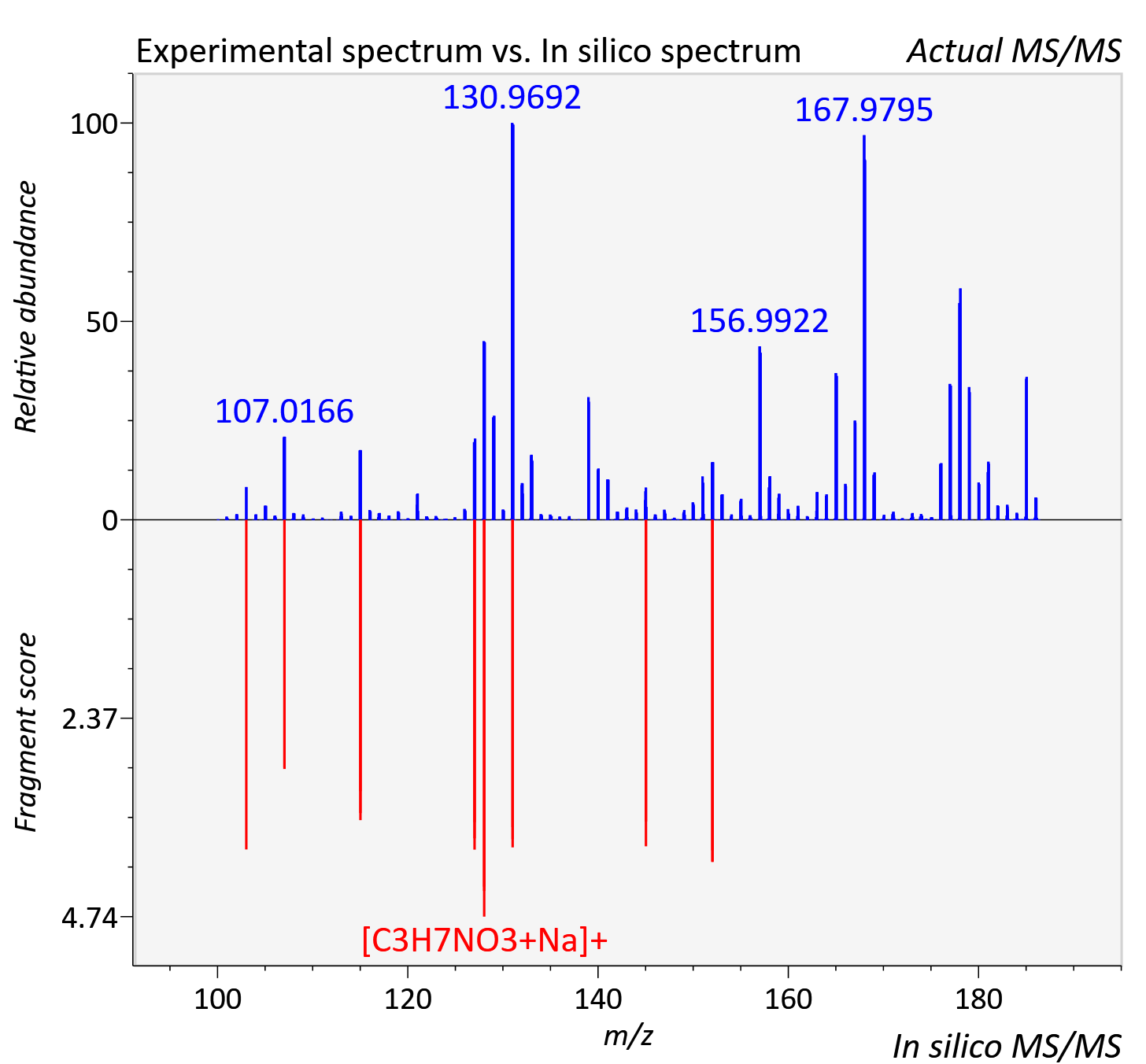
Experimental data
| Retention time | 7.77 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 186.038 | Theoretical mz | 186.037 |
| Error | 4.2 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.8717 |
Identifiers and class information
| Inchi key | NAALTSITEQTADZ-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(O)CNC(C(=O)O)CO |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| P08473 | MME | Neprilysin | T05409 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| Q9Y4C1 | KDM3A | Lysine-specific demethylase 3A | T25362 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| Q92820 | GGH | Gamma-glutamyl hydrolase | T71646 | SEA |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| Q9Y4G9 | PCSK6 | Proprotein convertase subtilisin/kexin type 6 | T86503 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T05409 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08473 | MME |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
