Compound details
(+)-Linderatin
| Compound ID | CDAMM01414 |
|---|---|
| Common name | (+)-Linderatin | IUPAC name | 3-phenyl-1-[2,4,6-trihydroxy-3-(3-methyl-6-propan-2-ylcyclohex-2-en-1-yl)phenyl]propan-1-one |
| Molecular formula | C25H30O4 |
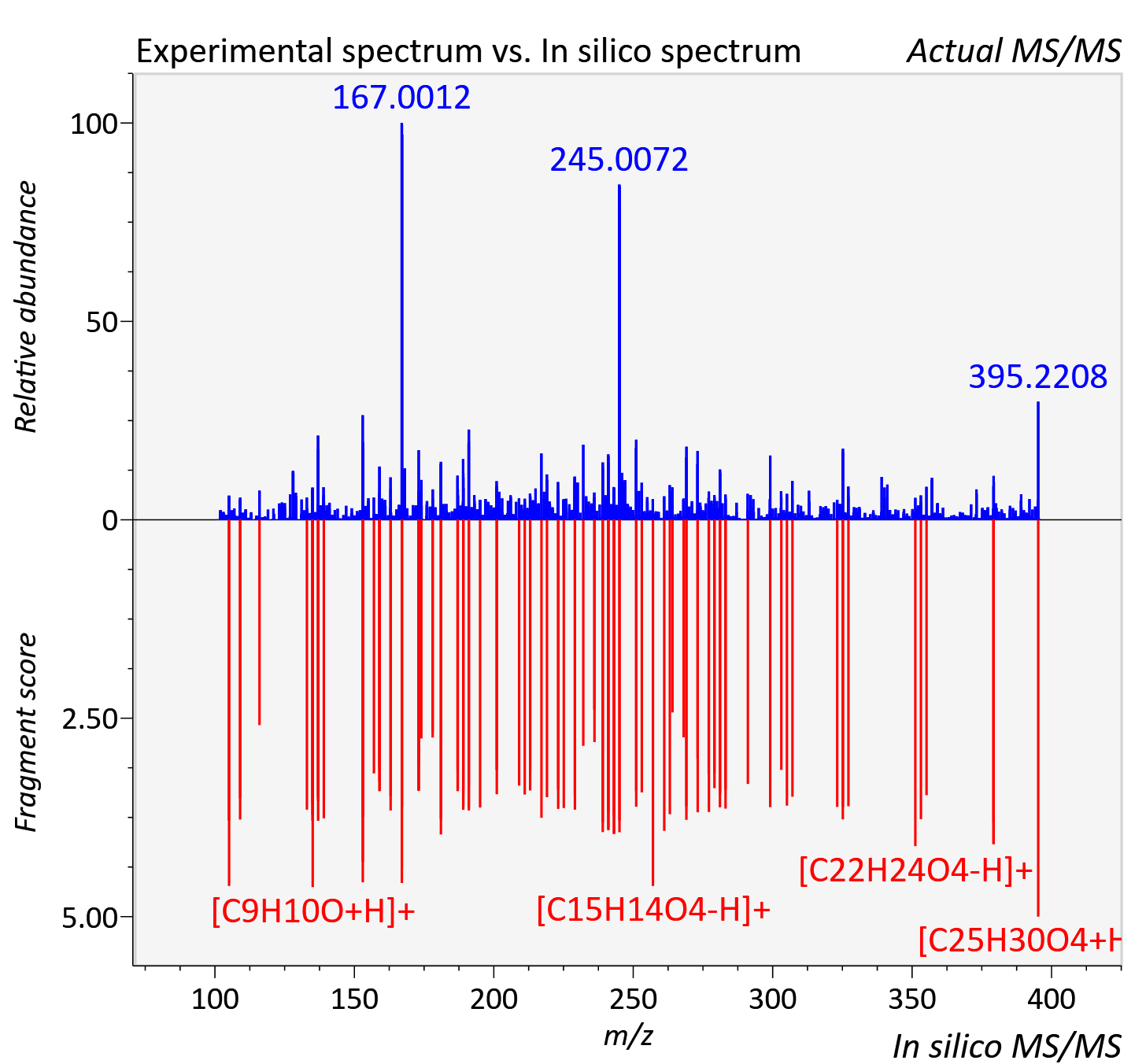
Experimental data
| Retention time | 4.98 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 395.22 | Theoretical mz | 395.221 |
| Error | 4.49 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0642 |
Identifiers and class information
| Inchi key | GHISAUFWVUOBIR-QINVSXPYNA-N |
|---|---|
| Smiles | O=C(C1=C(O)C=C(O)C(=C1O)C2C=C(C)CCC2C(C)C)CCC=3C=CC=CC3 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Linear 1,3-diarylpropanoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SwissTargetPrediction |
| P07195 | LDHB | L-lactate dehydrogenase B chain | T88057 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P07711 | CTSL | Cathepsin L | T98691 | SwissTargetPrediction |
| Q16548 | BCL2A1 | Bcl-2-related protein A1 (by homology) | T12797 | SEA |
| Q14330 | GPR18 | N-arachidonyl glycine receptor | T72267 | SEA |
| Q7RTX1 | TAS1R1 | Taste receptor | T41263 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T98691 | DI0263 | Middle East Respiratory Syndrome | [ICD-11: 1D64] | P07711 | CTSL |
| T98691 | DI0376 | Severe acute respiratory syndrome | [ICD-11: 1D65] | P07711 | CTSL |
| T41263 | DI0078 | Cholera | [ICD-11: 1A00] | Q7RTX1 | TAS1R1 |
