Compound details
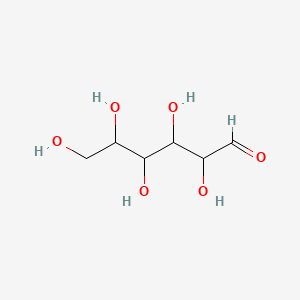
Glycoprotein-phospho-D-mannose
| Compound ID | CDAMM01384 |
|---|---|
| Common name | Glycoprotein-phospho-D-mannose | IUPAC name | 2,3,4,5,6-pentahydroxyhexanal |
| Molecular formula | C6H12O6 |
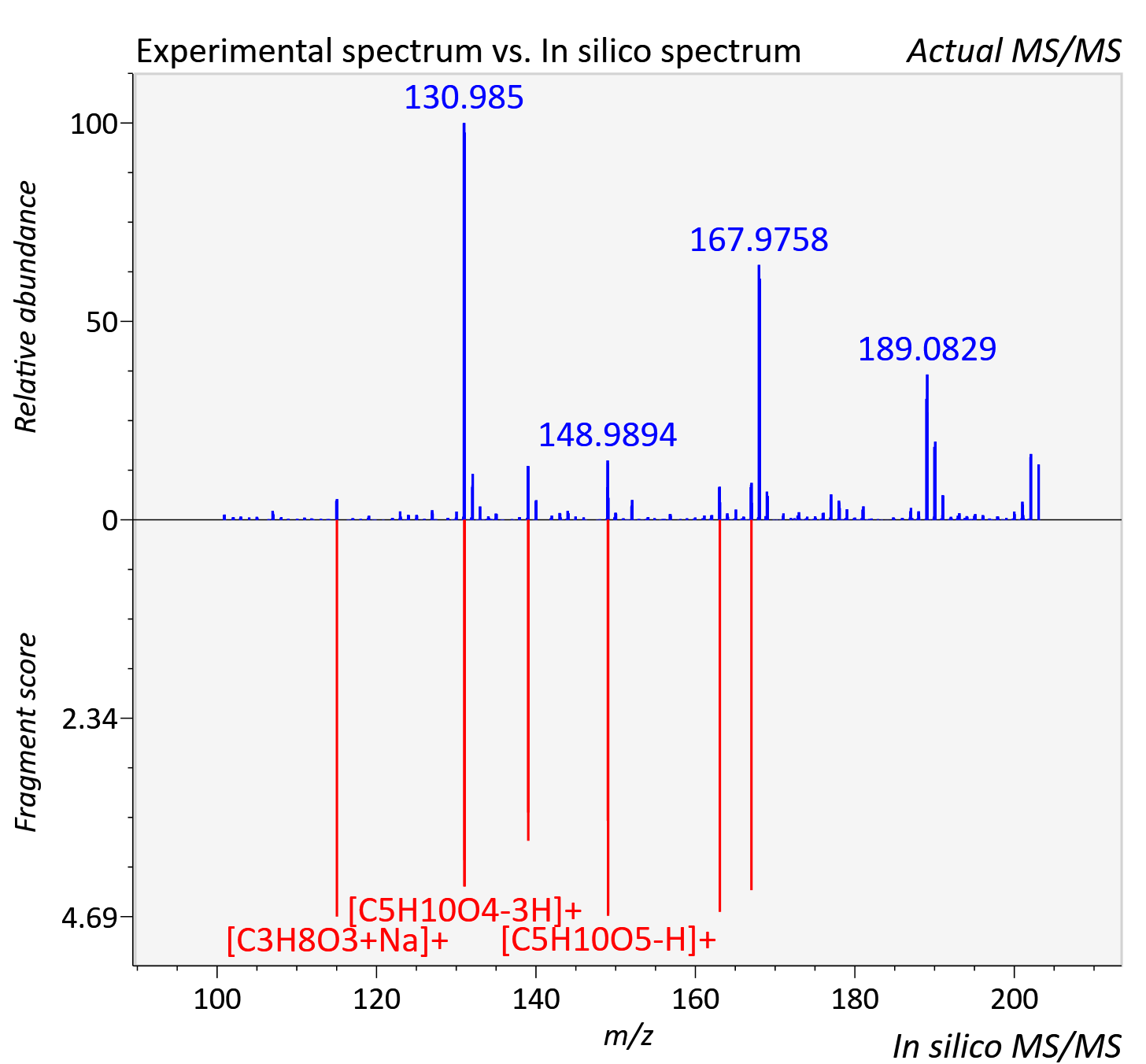
Experimental data
| Retention time | 9.91 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 203.05 | Theoretical mz | 203.052 |
| Error | 9.94 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.4656 |
Identifiers and class information
| Inchi key | GZCGUPFRVQAUEE-KVTDHHQDSA-N |
|---|---|
| Smiles | O=CC(O)C(O)C(O)C(O)CO |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SwissTargetPrediction |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SwissTargetPrediction |
| Q9UHC9 | NPC1L1 | Niemann-Pick C1-like protein 1 | T33901 | SwissTargetPrediction |
| P43116 | PTGER2 | Prostanoid EP2 receptor | T38529 | SwissTargetPrediction |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SwissTargetPrediction |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SwissTargetPrediction |
| P04062 | GBA | Beta-glucocerebrosidase | T84173 | SwissTargetPrediction |
| P43088 | PTGFR | Prostanoid FP receptor | T75797 | SwissTargetPrediction |
| P53396 | ACLY | ATP-citrate synthase | T52189 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T33901 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | Q9UHC9 | NPC1L1 |
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T84173 | DI0257 | Metabolic disorder | [ICD-11: 5C50-5D2Z] | P04062 | GBA |
| T75797 | DI0003 | Abortion | [ICD-11: JA00] | P43088 | PTGFR |
| T75797 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43088 | PTGFR |
| T75797 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43088 | PTGFR |
| T52189 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P53396 | ACLY |
