Compound details
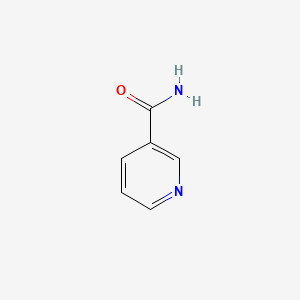
Niacinamide
| Compound ID | CDAMM01365 |
|---|---|
| Common name | Niacinamide | IUPAC name | pyridine-3-carboxamide |
| Molecular formula | C6H6N2O |
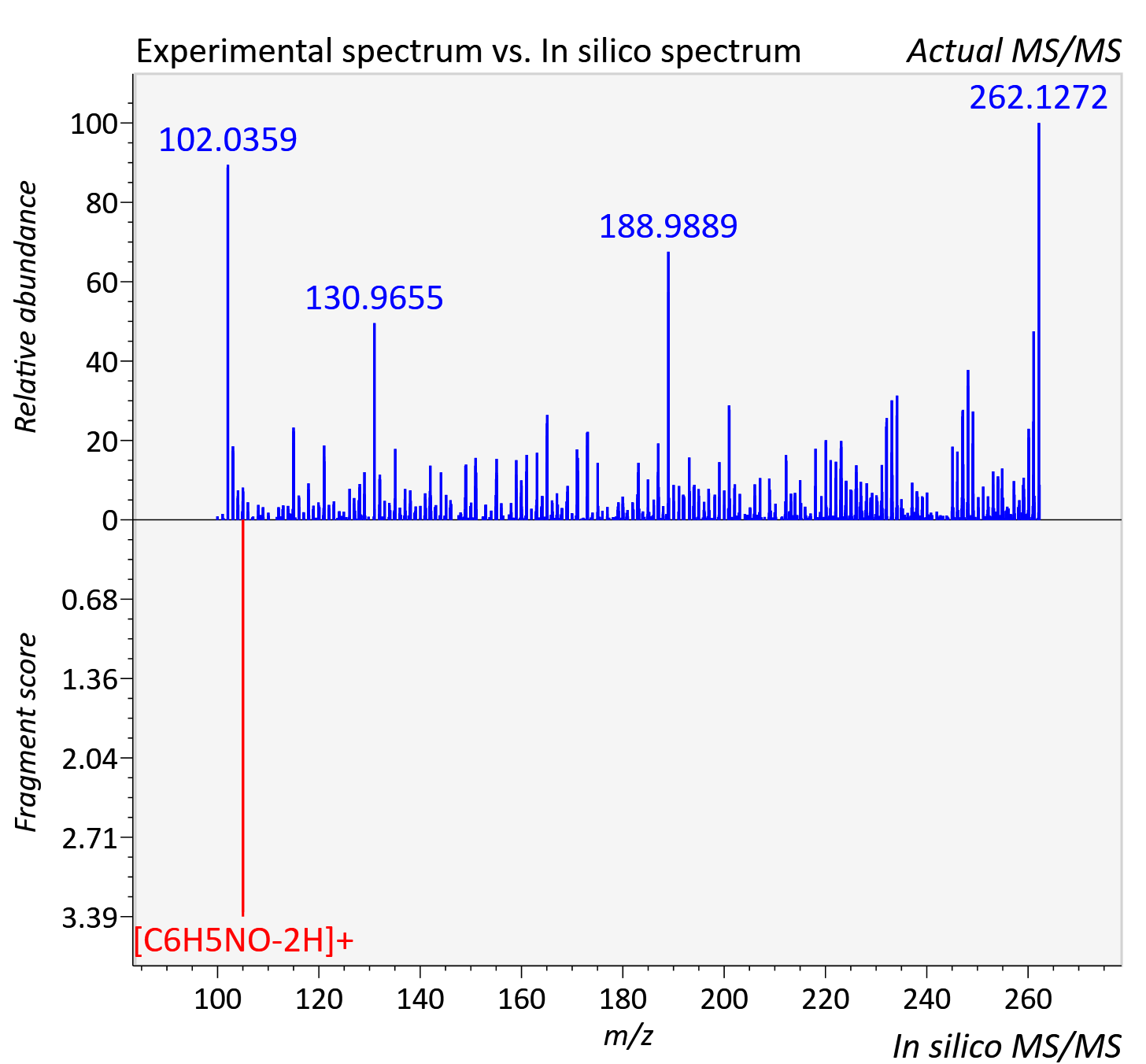
Experimental data
| Retention time | 14.04 |
|---|---|
| Adduct | [2M+NH4]+ |
| Actual mz | 262.128 | Theoretical mz | 262.13 |
| Error | 7.34 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.6625 |
Identifiers and class information
| Inchi key | DFPAKSUCGFBDDF-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(N)C=1C=NC=CC1 |
| Superclass | Organoheterocyclic compounds |
| Class | Pyridines and derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SEA |
| Q9NTG7 | SIRT3 | NAD-dependent deacetylase sirtuin 3 | T91959 | SwissTargetPrediction and SEA |
| Q8IXJ6 | SIRT2 | NAD-dependent deacetylase sirtuin 2 | T83904 | SwissTargetPrediction and SEA |
| P34913 | EPHX2 | Epoxide hydratase | T35734 | SEA |
| P11509 | CYP2A6 | Cytochrome P450 2A6 | T06455 | SEA |
| P15538 | CYP11B1 | Cytochrome P450 11B1 | T84621 | SEA |
| P19099 | CYP11B2 | Cytochrome P450 11B2 | T59056 | SEA |
| P21980 | TGM2 | Protein-glutamine gamma-glutamyltransferase | T05387 | SEA |
| Q92769 | HDAC2 | Histone deacetylase 2 | T51191 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SEA |
| Q9Y4C1 | KDM3A | Lysine-specific demethylase 3A | T25362 | SEA |
| P21731 | TBXA2R | Thromboxane A2 receptor | T76198 | SEA |
| P25105 | PTAFR | Platelet activating factor receptor | T87023 | SEA |
| P05093 | CYP17A1 | Cytochrome P450 17A1 | T89041 | SEA |
| O96017 | CHEK2 | Serine/threonine-protein kinase Chk2 | T28713 | SEA |
| O00311 | CDC7 | CDC7/DBF4 (Cell division cycle 7-related protein kinase/Activator of S phase kinase) | T81358 | SEA |
| P07333 | CSF1R | Macrophage colony stimulating factor receptor | T20333 | SEA |
| P49336 | CDK8 | Cell division protein kinase 8 | T10822 | SEA |
| P51686 | CCR9 | C-C chemokine receptor type 9 | T97873 | SEA |
| P43490 | NAMPT | Nicotinamide phosphoribosyltransferase | T54582 | SEA |
| P51812 | RPS6KA3 | Ribosomal protein S6 kinase alpha 3 | T03279 | SEA |
| P01130 | LDLR | LDL receptor | T94692 | SEA |
| O95819 | MAP4K4 | Mitogen-activated protein kinase kinase kinase kinase 4 | T71367 | SEA |
| P29323 | EPHB2 | Ephrin type-B receptor 2 | T73756 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P40337 | VHL | Von Hippel-Lindau disease tumor suppressor/Elongin B/Elongin C | T80423 | SEA |
| P42681 | TXK | Tyrosine-protein kinase TXK | T32590 | SEA |
| Q9Y337 | KLK5 | Kallikrein 5 | T11808 | SEA |
| P30532 | CHRNA5 | Nicotinic acetylcholine receptor alpha4/beta2/alpha5 | T00299 | SEA |
| P17787 | CHRNB2 | Nicotinic acetylcholine receptor alpha2/beta2 | T82543 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P24557 | TBXAS1 | Thromboxane-A synthase | T78356 | SEA |
| P13674 | P4HA1 | Prolyl 4-hydroxylasesubunit alpha-1 | T61729 | SEA |
| Q9P0G3 | KLK14 | Kallikrein-14 | T39195 | SEA |
| P20813 | CYP2B6 | Cytochrome P450 2B6 | T11793 | SEA |
| P49862 | KLK7 | Kallikrein-7 | T79155 | SEA |
| Q16637 | SMN2 | Survival motor neuron protein | T71907 | SEA |
| Q9NR20 | DYRK4 | Dual-specificity tyrosine-phosphorylation regulated kinase 4 | T20542 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T83904 | DI0365 | Retinopathy | [ICD-11: 9B71] | Q8IXJ6 | SIRT2 |
| T35734 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P34913 | EPHX2 |
| T35734 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P34913 | EPHX2 |
| T06455 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P11509 | CYP2A6 |
| T06455 | DI0351 | Psoriasis | [ICD-11: EA90] | P11509 | CYP2A6 |
| T84621 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P15538 | CYP11B1 |
| T84621 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P15538 | CYP11B1 |
| T59056 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P19099 | CYP11B2 |
| T05387 | DI0092 | Coeliac disease | [ICD-11: DA95] | P21980 | TGM2 |
| T05387 | DI0302 | Non-alcoholic fatty liver disease | [ICD-11: DB92] | P21980 | TGM2 |
| T51191 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q92769 | HDAC2 |
| T51191 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q92769 | HDAC2 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T76198 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P21731 | TBXA2R |
| T76198 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P21731 | TBXA2R |
| T76198 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P21731 | TBXA2R |
| T76198 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P21731 | TBXA2R |
| T87023 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P25105 | PTAFR |
| T89041 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P05093 | CYP17A1 |
| T28713 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O96017 | CHEK2 |
| T81358 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | O00311 | CDC7 |
| T81358 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O00311 | CDC7 |
| T20333 | DI0058 | Bone/articular cartilage neoplasm | [ICD-11: 2F7B] | P07333 | CSF1R |
| T10822 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P49336 | CDK8 |
| T10822 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P49336 | CDK8 |
| T97873 | DI0107 | Crohn disease | [ICD-11: DD70] | P51686 | CCR9 |
| T54582 | DI0231 | Leukaemia | [ICD-11: 2A60-2B33] | P43490 | NAMPT |
| T54582 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P43490 | NAMPT |
| T54582 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P43490 | NAMPT |
| T03279 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | P51812 | RPS6KA3 |
| T94692 | DI0238 | Lung cancer | [ICD-11: 2C25] | P01130 | LDLR |
| T73756 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P29323 | EPHB2 |
| T73756 | DI0252 | Melanoma | [ICD-11: 2C30] | P29323 | EPHB2 |
| T73756 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | P29323 | EPHB2 |
| T73756 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P29323 | EPHB2 |
| T82543 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P17787 | CHRNB2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T78356 | DI0030 | Angina pectoris | [ICD-11: BA40] | P24557 | TBXAS1 |
| T61729 | DI0178 | Hepatic fibrosis/cirrhosis | [ICD-11: DB93] | P13674 | P4HA1 |
| T61729 | DI0413 | Transplant rejection | [ICD-11: NE84] | P13674 | P4HA1 |
| T71907 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | Q16637 | SMN2 |
