Compound details
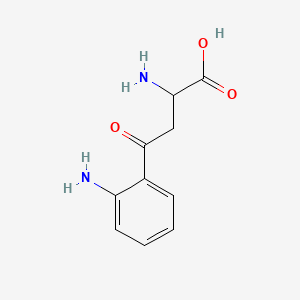
L-Kynurenine
| Compound ID | CDAMM01362 |
|---|---|
| Common name | L-Kynurenine | IUPAC name | 2-amino-4-(2-aminophenyl)-4-oxobutanoic acid |
| Molecular formula | C10H12N2O3 |
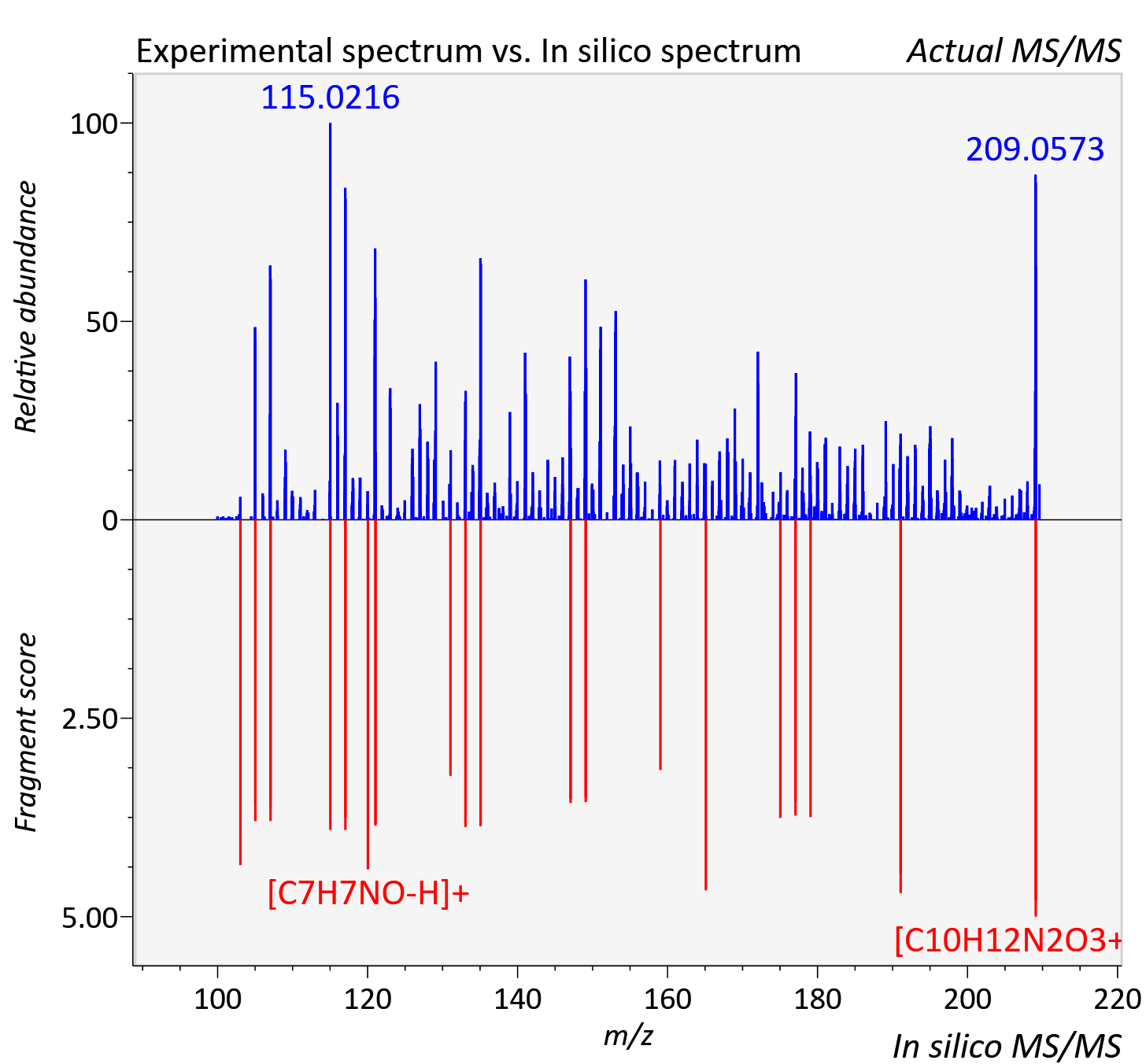
Experimental data
| Retention time | 8.15 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 209.091 | Theoretical mz | 209.092 |
| Error | 4.19 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5837 |
Identifiers and class information
| Inchi key | YGPSJZOEDVAXAB-QMMMGPOBSA-N |
|---|---|
| Smiles | O=C(O)C(N)CC(=O)C=1C=CC=CC1N |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P15144 | ANPEP | Aminopeptidase N | T67272 | SEA |
| P09960 | LTA4H | Leukotriene A4 hydrolase | T03691 | SEA |
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P43005 | SLC1A1 | Excitatory amino acid transporter 3 | T31721 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| Q13002 | GRIK2 | Glutamate receptor ionotropic kainate 2 | T58178 | SEA |
| Q14832 | GRM3 | Metabotropic glutamate receptor 3 | T02719 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q16478 | GRIK5 | Glutamate receptor ionotropic kainate 5 | T87930 | SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| P42262 | GRIA2 | Glutamate receptor ionotropic, AMPA 2 | T42392 | SEA |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SEA |
| O15229 | KMO | Kynurenine 3-monooxygenase | T50973 | SEA |
| P07101 | TH | Tyrosine 3-hydroxylase | T62390 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SwissTargetPrediction and SEA |
| Q01650 | SLC7A5 | L-type amino acid transporter 1 (by homology) | T48330 | SEA |
| P52732 | KIF11 | Kinesin-like protein 1 | T28484 | SEA |
| Q9ULA0 | DNPEP | Aspartyl aminopeptidase | T45287 | SEA |
| P15085 | CPA1 | Carboxypeptidase A1 | T33966 | SEA |
| P43003 | SLC1A3 | Excitatory amino acid transporter 1 | T86582 | SEA |
| P05089 | ARG1 | Arginase-1 (by homology) | T63224 | SEA |
| Q9Y256 | RCE1 | Prenyl protein specific protease | T10756 | SEA |
| Q03405 | PLAUR | Urokinase plasminogen activator surface receptor | T74363 | SEA |
| P15169 | CPN1 | Carboxypeptidase N, catalytic subunit | T34471 | SEA |
| O60235 | TMPRSS11D | Transmembrane protease serine 11D | T00302 | SEA |
| Q8IWU9 | TPH2 | Tryptophan hydroxylase | T51593 | SEA |
| Q9UPY5 | SLC7A11 | Cystine/glutamate transporter | T11615 | SEA |
| Q9BWQ3 | ASCT2 | Alanine/serine/cysteine transporter 2 | T59493 | SEA |
| P42263 | GRIA3 | Glutamate receptor AMPA 3 | T62391 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T67272 | DI0351 | Psoriasis | [ICD-11: EA90] | P15144 | ANPEP |
| T03691 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P09960 | LTA4H |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T02719 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q14832 | GRM3 |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T42392 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42262 | GRIA2 |
| T62390 | DI0019 | Adrenomedullary hyperfunction | [ICD-11: 5A75] | P07101 | TH |
| T62390 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P07101 | TH |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T28484 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P52732 | KIF11 |
| T28484 | DI0172 | Head and neck cancer | [ICD-11: 2D42] | P52732 | KIF11 |
| T28484 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P52732 | KIF11 |
| T28484 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P52732 | KIF11 |
| T28484 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | P52732 | KIF11 |
| T28484 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P52732 | KIF11 |
| T28484 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P52732 | KIF11 |
| T28484 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P52732 | KIF11 |
| T45287 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q9ULA0 | DNPEP |
| T63224 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P05089 | ARG1 |
| T74363 | DI0286 | Myeloproliferative neoplasm | [ICD-11: 2A20] | Q03405 | PLAUR |
| T74363 | DI0404 | Thrombocytosis | [ICD-11: 3B63] | Q03405 | PLAUR |
| T51593 | DI0066 | Carcinoid syndrome | [ICD-11: 5B10] | Q8IWU9 | TPH2 |
| T51593 | DI0389 | Small intestine developmental anomaly | [ICD-11: DA90] | Q8IWU9 | TPH2 |
| T11615 | DI0054 | Body-focused behaviour disorder | [ICD-11: 6B25] | Q9UPY5 | SLC7A11 |
| T62391 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42263 | GRIA3 |
