Compound details
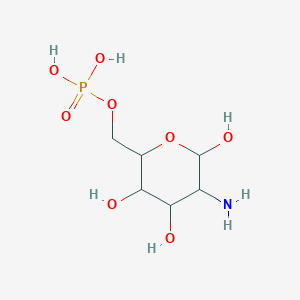
Glucosamine 6-phosphate
| Compound ID | CDAMM01361 |
|---|---|
| Common name | Glucosamine 6-phosphate | IUPAC name | (5-amino-3,4,6-trihydroxyoxan-2-yl)methyl dihydrogen phosphate |
| Molecular formula | C6H14NO8P |
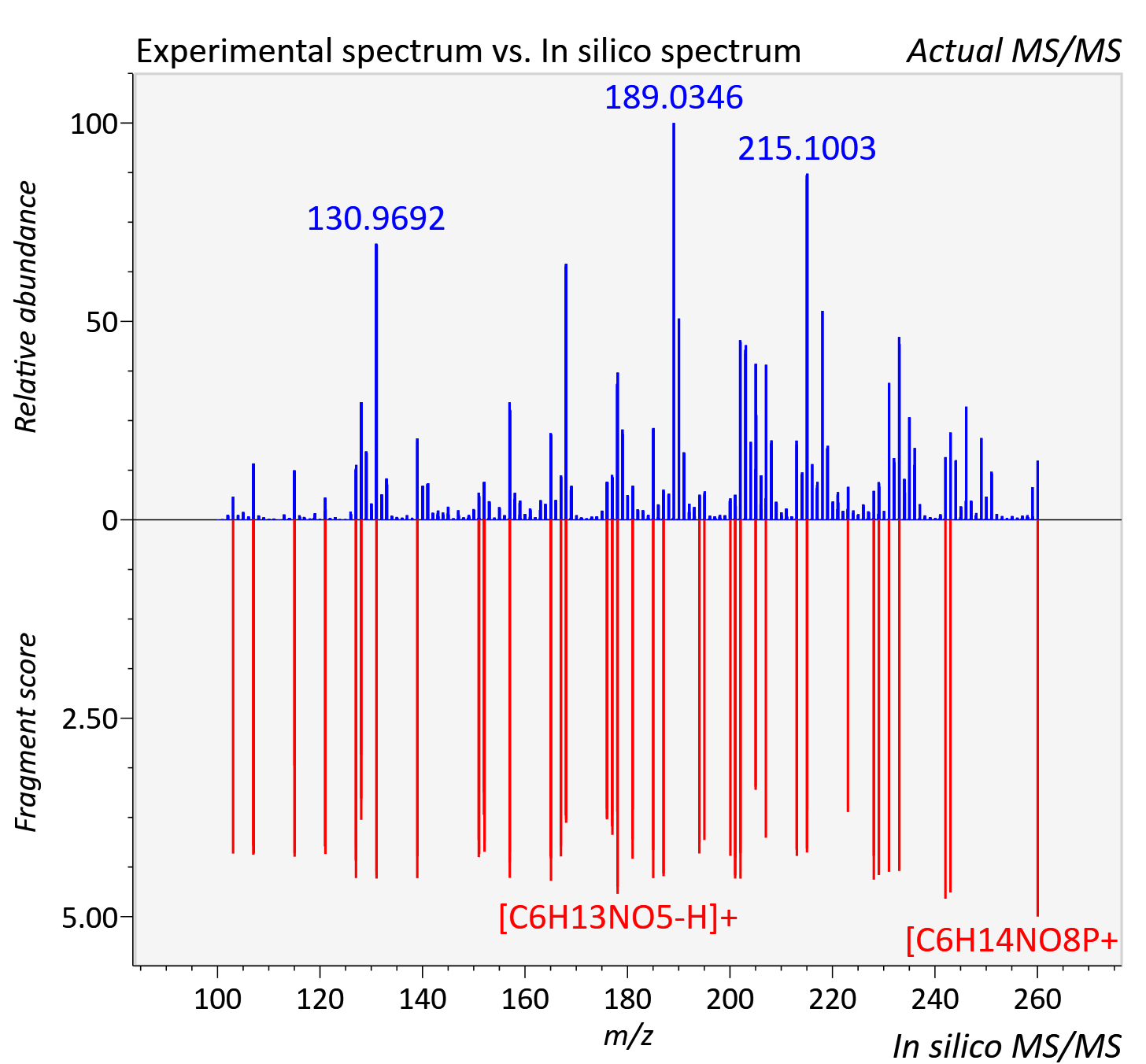
Experimental data
| Retention time | 7.77 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 260.051 | Theoretical mz | 260.053 |
| Error | 9.52 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.9637 |
Identifiers and class information
| Inchi key | XHMJOUIAFHJHBW-UKFBFLRUSA-N |
|---|---|
| Smiles | O=P(O)(O)OCC1OC(O)C(N)C(O)C1O |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P04818 | TYMS | Thymidylate synthase (by homology) | T98397 | SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
| P47900 | P2RY1 | Purinergic receptor P2Y1 | T67818 | SEA |
| P41231 | P2RY2 | Purinergic receptor P2Y2 | T93515 | SEA |
| Q15077 | P2RY6 | Pyrimidinergic receptor P2Y6 | T90782 | SEA |
| P63000 | RAC1 | Ras-related C3 botulinum toxin substrate 1 | T88752 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| P11172 | UMPS | Uridine 5'-monophosphate synthase | T15851 | SEA |
| O94759 | TRPM2 | Transient receptor potential cation channel subfamily M member 2 | T78205 | SEA |
| Q15391 | P2RY14 | Purinergic receptor P2Y14 | T16555 | SEA |
| Q05823 | RNASEL | RNase L | T25200 | SEA |
| P51575 | P2RX1 | P2X purinoceptor 1 | T69091 | SEA |
| Q14643 | ITPR1 | Inositol 1,4,5-trisphosphate receptor type 1 | T82624 | SEA |
| Q14643 | ITPR1 | Inositol 1,4,5-trisphosphate receptor type 1 | T82624 | SEA |
| Q99571 | P2RX4 | P2X purinoceptor 4 | T60330 | SEA |
| P11717 | IGF2R | Insulin-like growth factor II receptor | T59550 | SEA |
| Q13304 | GPR17 | Uracil nucleotide/cysteinyl leukotriene receptor | T33902 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T98397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P04818 | TYMS |
| T98397 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P04818 | TYMS |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
| T93515 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P41231 | P2RY2 |
| T90782 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15077 | P2RY6 |
| T88752 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P63000 | RAC1 |
| T82624 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q14643 | ITPR1 |
| T82624 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q14643 | ITPR1 |
| T59550 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | P11717 | IGF2R |
| T59550 | DI0368 | Sarcoidosis | [ICD-11: 4B20] | P11717 | IGF2R |
