Compound details
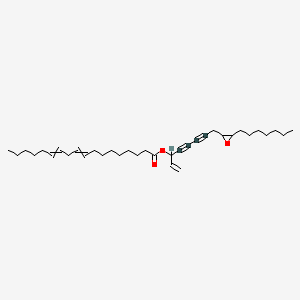
Panaxydol linoleate
| Compound ID | CDAMM01330 |
|---|---|
| Common name | Panaxydol linoleate | IUPAC name | 8-(3-heptyloxiran-2-yl)oct-1-en-4,6-diyn-3-yl octadeca-9,12-dienoate |
| Molecular formula | C35H54O3 |
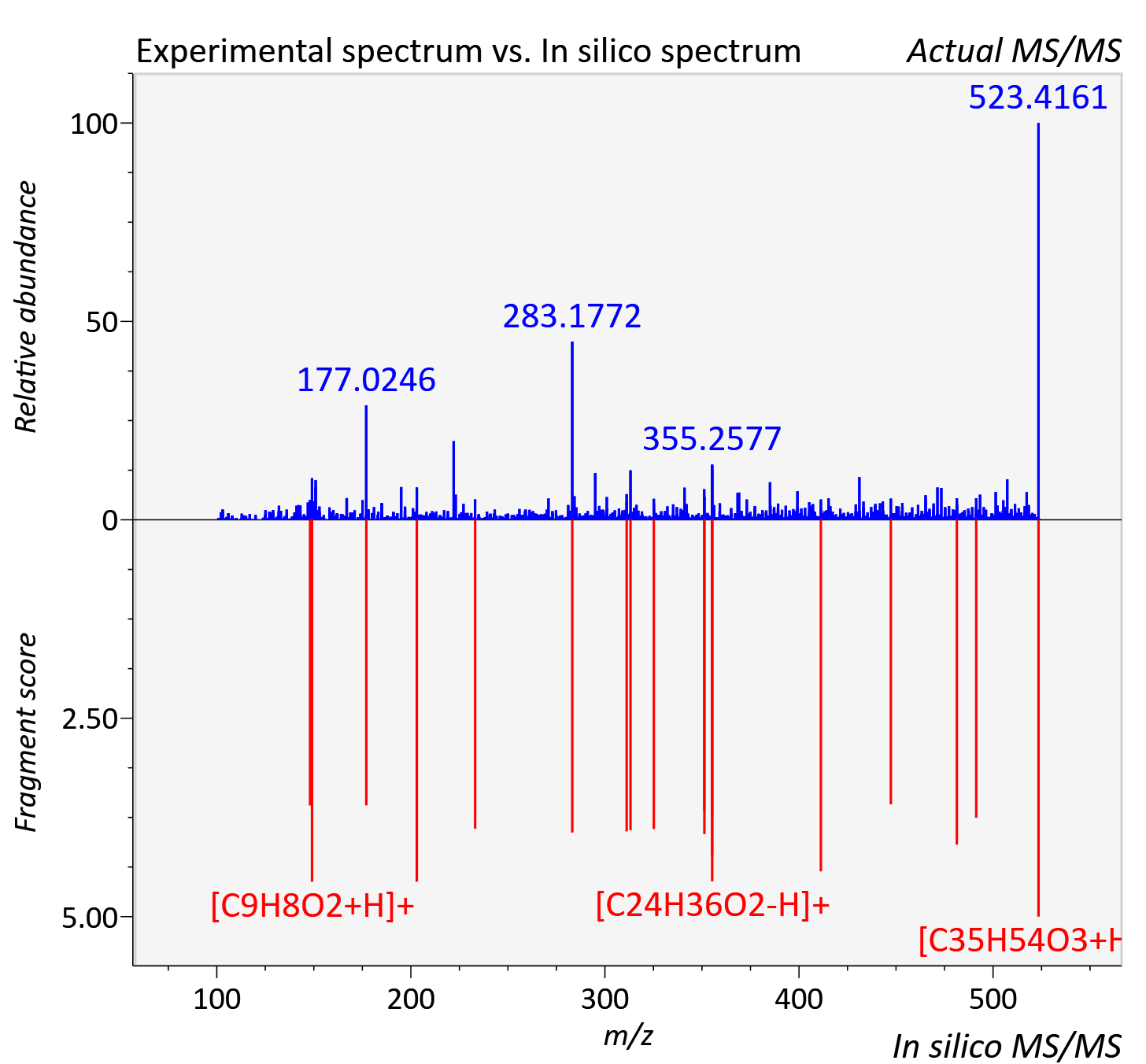
Experimental data
| Retention time | 19.33 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 523.414 | Theoretical mz | 523.414 |
| Error | 0.15 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.2382 |
Identifiers and class information
| Inchi key | MGQITZGZKLABCX-XUPYMNJSNA-N |
|---|---|
| Smiles | O=C(OC(C#CC#CCC1OC1CCCCCCC)C=C)CCCCCCCC=CCC=CCCCCC |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SEA |
| Q02156 | PRKCE | Protein kinase C epsilon | T00895 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9Y4D2 | DAGLA | Sn1-specific diacylglycerol lipase alpha | T03150 | SEA |
| Q13822 | ENPP2 | Autotaxin | T63512 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| Q9UP65 | PLA2G4C | Cytosolic phospholipase A2 gamma | T91113 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T00895 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | Q02156 | PRKCE |
| T00895 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q02156 | PRKCE |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T63512 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q13822 | ENPP2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
