Compound details
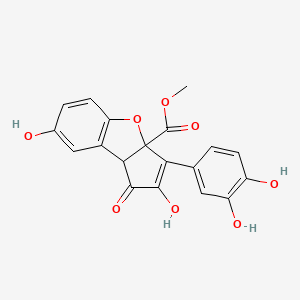
Torosaflavone D
| Compound ID | CDAMM01285 |
|---|---|
| Common name | Torosaflavone D | IUPAC name | methyl 3-(3,4-dihydroxyphenyl)-2,7-dihydroxy-1-oxo-8bH-cyclopenta[b][1]benzofuran-3a-carboxylate |
| Molecular formula | C19H14O8 |
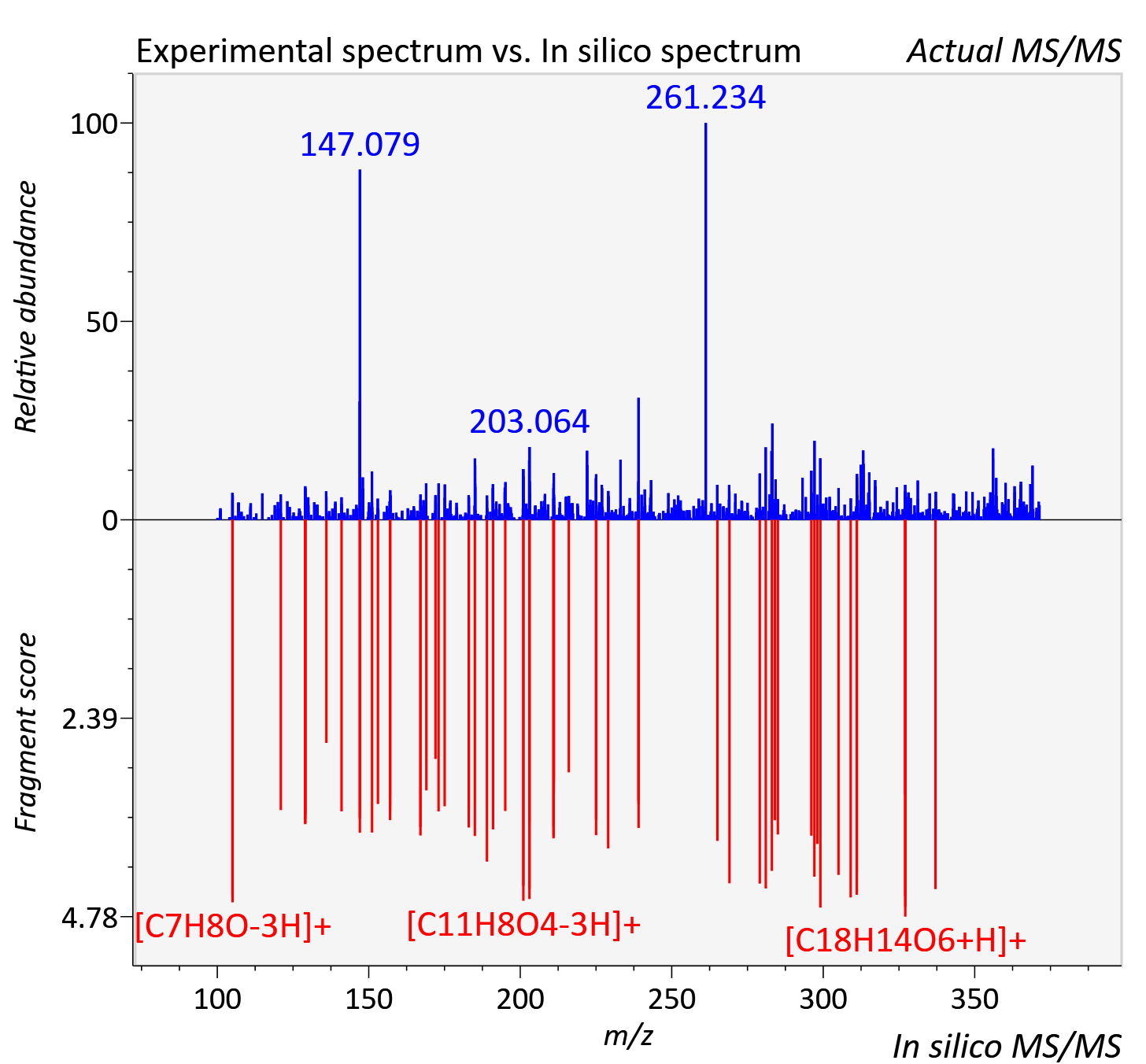
Experimental data
| Retention time | 21.57 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 371.076 | Theoretical mz | 371.076 |
| Error | 0.06 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.6911 |
Identifiers and class information
| Inchi key | YGQFRAJVEBXHHA-HWKANZROSA-N |
|---|---|
| Smiles | O=C(OC)C12OC3=CC=C(O)C=C3C2C(=O)C(O)=C1C=4C=CC(O)=C(O)C4 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SwissTargetPrediction |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SwissTargetPrediction |
| P05121 | SERPINE1 | Plasminogen activator inhibitor-1 | T15556 | SwissTargetPrediction |
| O14757 | CHEK1 | Serine/threonine-protein kinase Chk1 | T62449 | SwissTargetPrediction |
| P14061 | HSD17B1 | Estradiol 17-beta-dehydrogenase 1 | T44011 | SwissTargetPrediction |
| P30291 | WEE1 | Serine/threonine-protein kinase WEE1 | T69991 | SwissTargetPrediction |
| P11802 | CDK4 | Cyclin-dependent kinase 4 | T85799 | SwissTargetPrediction |
| P56817 | BACE1 | Beta-secretase 1 | T79031 | SwissTargetPrediction |
| Q9HBH9 | MKNK2 | MAP kinase signal-integrating kinase 2 | T99160 | SwissTargetPrediction |
| Q9NPH5 | NOX4 | NADPH oxidase 4 | T29741 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| Q9HC98 | NEK6 | Serine/threonine-protein kinase NEK6 | T78992 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T15556 | DI0037 | Asthma | [ICD-11: CA23] | P05121 | SERPINE1 |
| T15556 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P05121 | SERPINE1 |
| T62449 | DI0027 | Anal cancer | [ICD-11: 2C00] | O14757 | CHEK1 |
| T62449 | DI0172 | Head and neck cancer | [ICD-11: 2D42] | O14757 | CHEK1 |
| T62449 | DI0183 | Hodgkin lymphoma | [ICD-11: 2B30] | O14757 | CHEK1 |
| T62449 | DI0238 | Lung cancer | [ICD-11: 2C25] | O14757 | CHEK1 |
| T62449 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | O14757 | CHEK1 |
| T62449 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | O14757 | CHEK1 |
| T62449 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | O14757 | CHEK1 |
| T62449 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O14757 | CHEK1 |
| T69991 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P30291 | WEE1 |
| T69991 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P30291 | WEE1 |
| T69991 | DI0429 | Uterine ligament/parametrium/uterine adnexa neoplasm | [ICD-11: 2C72] | P30291 | WEE1 |
| T85799 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11802 | CDK4 |
| T85799 | DI0238 | Lung cancer | [ICD-11: 2C25] | P11802 | CDK4 |
| T85799 | DI0351 | Psoriasis | [ICD-11: EA90] | P11802 | CDK4 |
| T79031 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P56817 | BACE1 |
| T99160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q9HBH9 | MKNK2 |
| T99160 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | Q9HBH9 | MKNK2 |
| T99160 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q9HBH9 | MKNK2 |
| T99160 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q9HBH9 | MKNK2 |
| T99160 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9HBH9 | MKNK2 |
| T29741 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9NPH5 | NOX4 |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
