Compound details
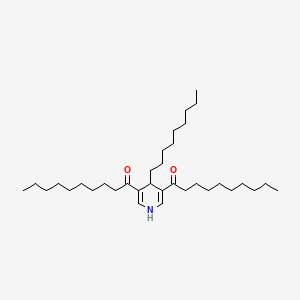
1,1\'-(1,4-Dihydro-4-nonyl-3,5-pyridinediyl)bis[1-decanone]
| Compound ID | CDAMM01281 |
|---|---|
| Common name | 1,1\'-(1,4-Dihydro-4-nonyl-3,5-pyridinediyl)bis[1-decanone] | IUPAC name | 1-(5-decanoyl-4-nonyl-1,4-dihydropyridin-3-yl)decan-1-one |
| Molecular formula | C34H61NO2 |
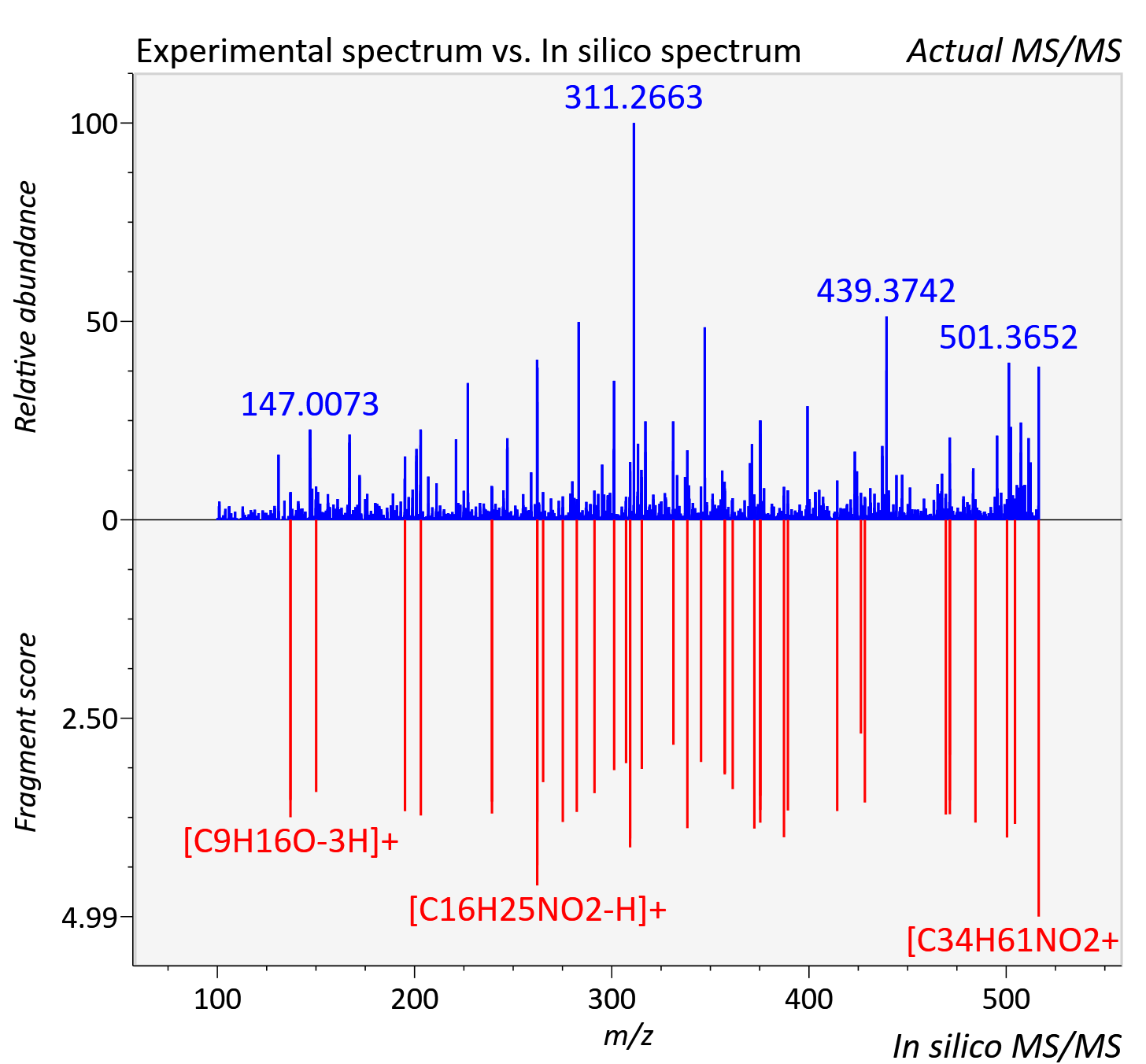
Experimental data
| Retention time | 15.18 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 516.475 | Theoretical mz | 516.477 |
| Error | 4.79 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.6408 |
Identifiers and class information
| Inchi key | WBCPDCHBBWAFSO-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(C1=CNC=C(C(=O)CCCCCCCCC)C1CCCCCCCCC)CCCCCCCCC |
| Superclass | Organoheterocyclic compounds |
| Class | Pyridines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P42336 | PIK3CA | PI3-kinase p110-alpha subunit | T80276 | SwissTargetPrediction |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SwissTargetPrediction and SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction |
| Q07869 | PPARA | Peroxisome proliferator-activated receptor alpha | T86591 | SwissTargetPrediction |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SEA |
| P10827 | THRA | Thyroid hormone receptor alpha | T79591 | SEA |
| P10828 | THRB | Thyroid hormone receptor beta-1 | T98933 | SEA |
| P43116 | PTGER2 | Prostanoid EP2 receptor | T38529 | SEA |
| O00329 | PIK3CD | PI3-kinase p110-delta subunit | T67849 | SwissTargetPrediction |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9Y4D2 | DAGLA | Sn1-specific diacylglycerol lipase alpha | T03150 | SwissTargetPrediction and SEA |
| Q9BV23 | ABHD6 | Monoacylglycerol lipase ABHD6 | T81452 | SwissTargetPrediction |
| Q13093 | PLA2G7 | LDL-associated phospholipase A2 | T69912 | SwissTargetPrediction |
| Q8NCG7 | DAGLB | Sn1-specific diacylglycerol lipase beta | T15509 | SwissTargetPrediction |
| Q9H228 | S1PR5 | Sphingosine 1-phosphate receptor Edg-8 | T50089 | SEA |
| Q13822 | ENPP2 | Autotaxin | T63512 | SEA |
| O60906 | SMPD2 | Neutral sphingomyelinase | T57316 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T80276 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P42336 | PIK3CA |
| T80276 | DI0151 | Follicular lymphoma | [ICD-11: 2A80] | P42336 | PIK3CA |
| T80276 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P42336 | PIK3CA |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T86591 | DI0187 | Hyperlipidemia | [ICD-11: 5C80] | Q07869 | PPARA |
| T86591 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | Q07869 | PPARA |
| T86591 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q07869 | PPARA |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T79591 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10827 | THRA |
| T79591 | DI0197 | Hypo-thyroidism | [ICD-11: 5A00] | P10827 | THRA |
| T98933 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10828 | THRB |
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
| T67849 | DI0151 | Follicular lymphoma | [ICD-11: 2A80] | O00329 | PIK3CD |
| T67849 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | O00329 | PIK3CD |
| T67849 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | O00329 | PIK3CD |
| T67849 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | O00329 | PIK3CD |
| T67849 | DI0395 | Stomach cancer | [ICD-11: 2B72] | O00329 | PIK3CD |
| T69912 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q13093 | PLA2G7 |
| T69912 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | Q13093 | PLA2G7 |
| T69912 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | Q13093 | PLA2G7 |
| T50089 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | Q9H228 | S1PR5 |
| T63512 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q13822 | ENPP2 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
