Compound details
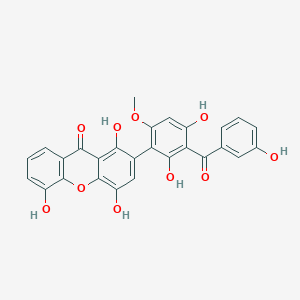
Garciduol B
| Compound ID | CDAMM01278 |
|---|---|
| Common name | Garciduol B | IUPAC name | 2-[2,4-dihydroxy-3-(3-hydroxybenzoyl)-6-methoxyphenyl]-1,4,5-trihydroxyxanthen-9-one |
| Molecular formula | C27H18O10 |
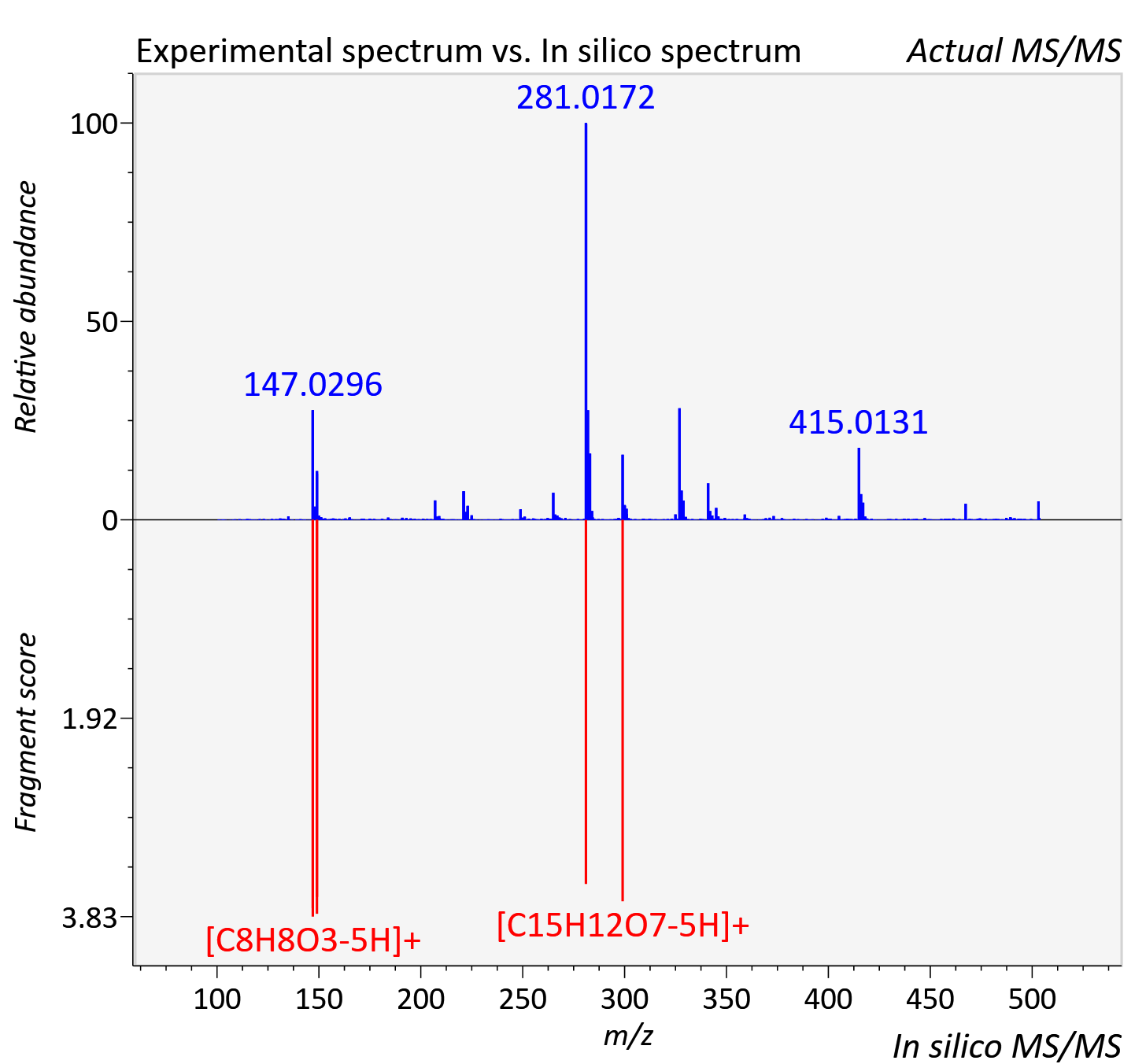
Experimental data
| Retention time | 17.53 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 503.092 | Theoretical mz | 503.097 |
| Error | 10.29 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7248 |
Identifiers and class information
| Inchi key | XKZFGJPNYONQOM-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(C=1C=CC=C(O)C1)C2=C(O)C=C(OC)C(=C2O)C=3C=C(O)C=4OC=5C(O)=CC=CC5C(=O)C4C3O |
| Superclass | Organoheterocyclic compounds |
| Class | Benzopyrans |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P21397 | MAOA | Monoamine oxidase A | T83875 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| O60911 | CTSV | Cathepsin (V and K) | T93653 | SEA |
| P32754 | HPD | 4-hydroxyphenylpyruvate dioxygenase | T07137 | SEA |
| O43826 | SLC37A4 | Glucose-6-phosphate translocase | T47306 | SEA |
| Q14721 | KCNB1 | Potassium voltage-gated channel subfamily B member 1 | T44516 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83875 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P21397 | MAOA |
| T83875 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P21397 | MAOA |
| T93653 | DI0055 | Bone cancer | [ICD-11: 2B5Z] | O60911 | CTSV |
| T93653 | DI0087 | Chronic pain | [ICD-11: MG30] | O60911 | CTSV |
| T07137 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P32754 | HPD |
