Compound details
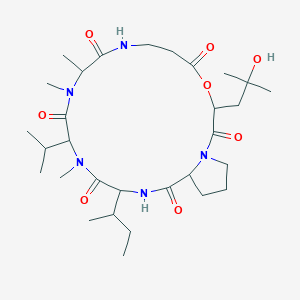
Hydroxydestruxin B
| Compound ID | CDAMM01255 |
|---|---|
| Common name | Hydroxydestruxin B | IUPAC name | 16-butan-2-yl-3-(2-hydroxy-2-methylpropyl)-10,11,14-trimethyl-13-propan-2-yl-4-oxa-1,8,11,14,17-pentazabicyclo[17.3.0]docosane-2,5,9,12,15,18-hexone |
| Molecular formula | C30H51N5O8 |
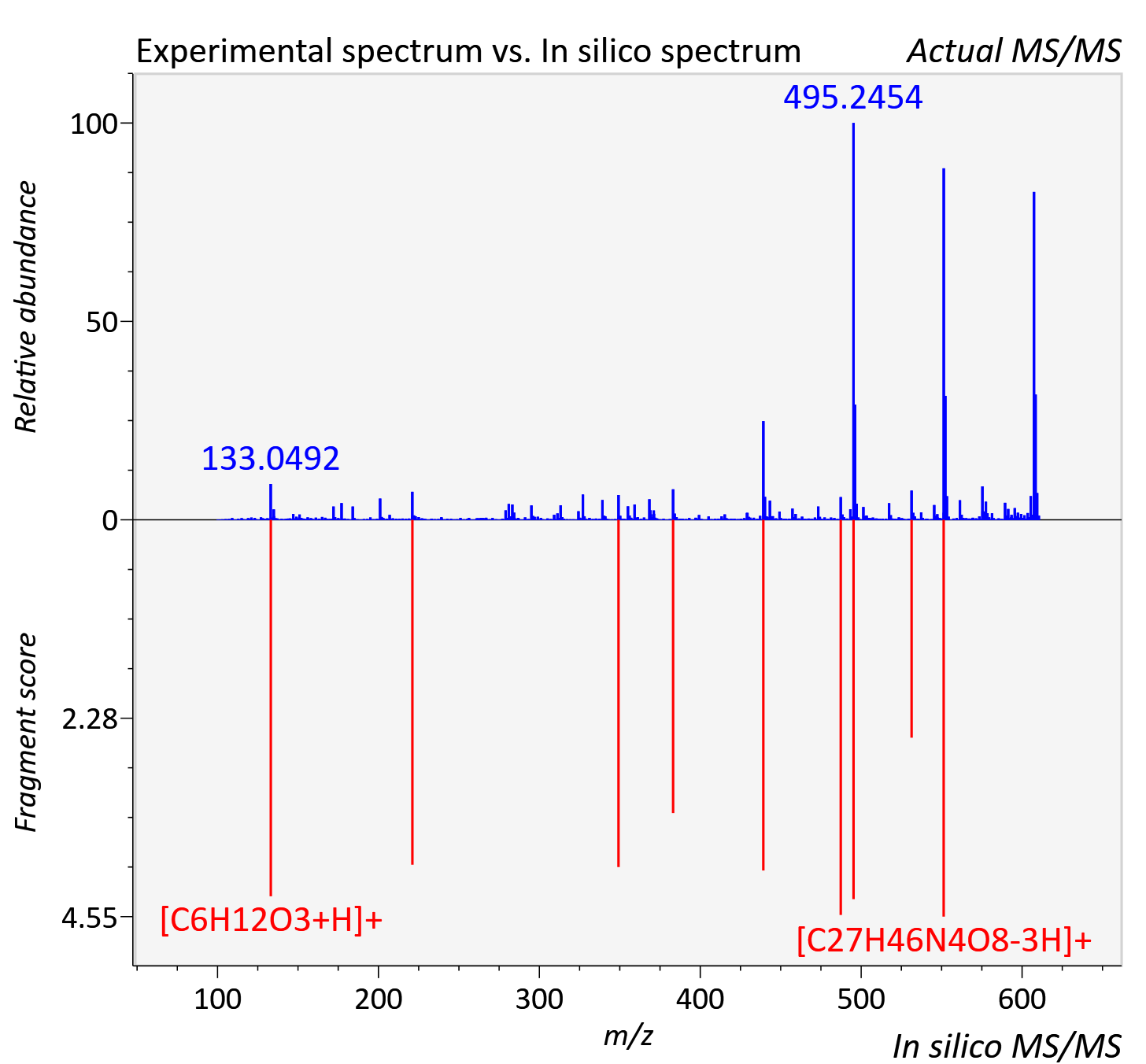
Experimental data
| Retention time | 20.31 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 610.388 | Theoretical mz | 610.381 |
| Error | 11.02 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0464 |
Identifiers and class information
| Inchi key | COCZASXGUADHMT-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(C(=O)N2CCCC2C(=O)NC(C(=O)N(C)C(C(=O)N(C)C(C(=O)NCC1)C)C(C)C)C(C)CC)CC(O)(C)C |
| Superclass | Organic acids and derivatives |
| Class | Peptidomimetics |
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T63816 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P56524 | HDAC4 |
| T06850 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | Q9UBU3 | GHRL |
| T59604 | DI0087 | Chronic pain | [ICD-11: MG30] | Q92847 | GHSR |
| T59604 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | Q92847 | GHSR |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
