Compound details
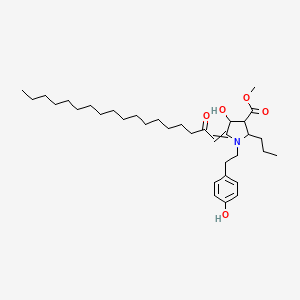
Plakoridine A
| Compound ID | CDAMM01209 |
|---|---|
| Common name | Plakoridine A | IUPAC name | methyl 4-hydroxy-1-[2-(4-hydroxyphenyl)ethyl]-5-(2-oxooctadecylidene)-2-propylpyrrolidine-3-carboxylate |
| Molecular formula | C35H57NO5 |
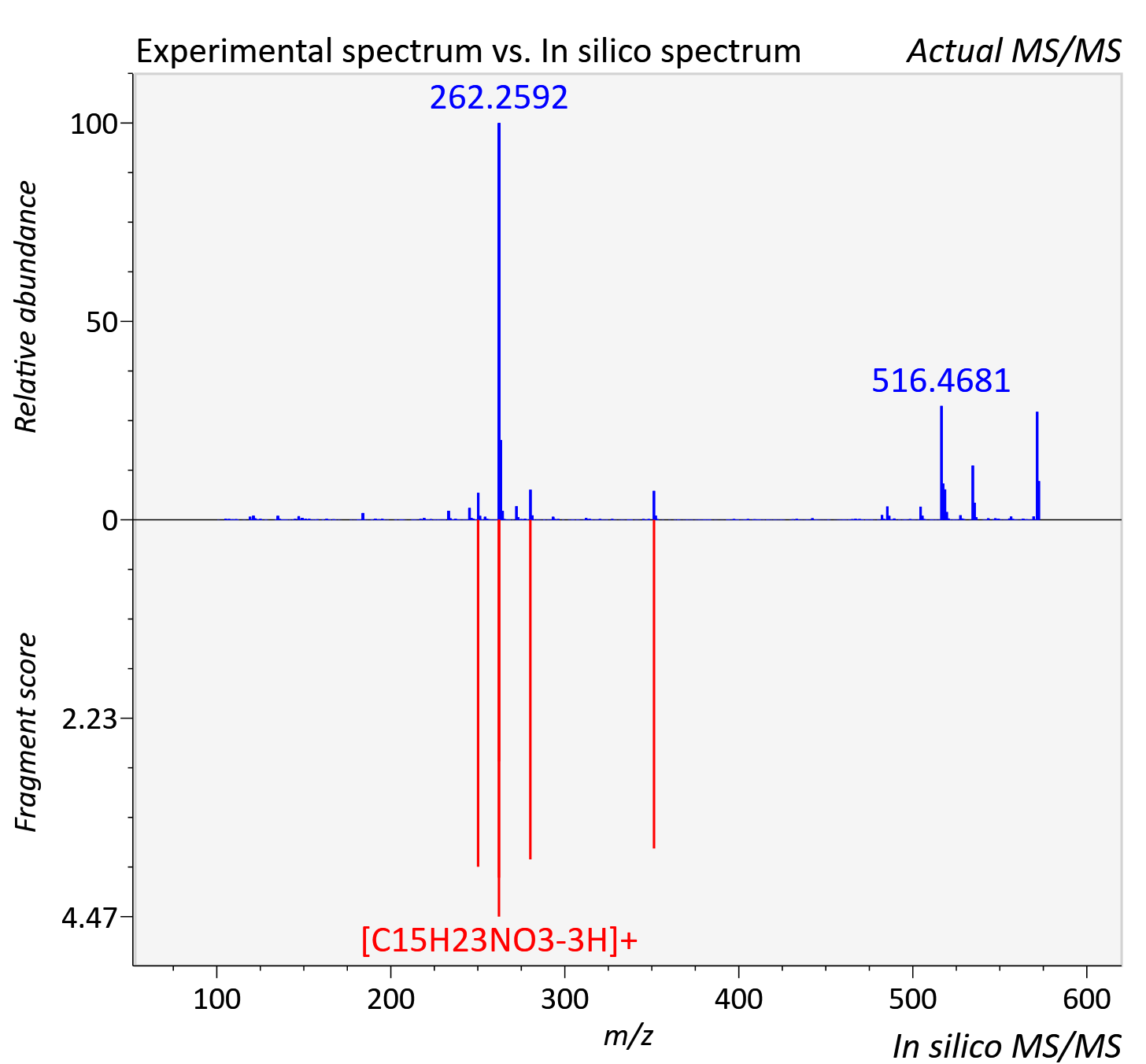
Experimental data
| Retention time | 16.01 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 572.434 | Theoretical mz | 572.431 |
| Error | 4.93 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.6054 |
Identifiers and class information
| Inchi key | ULFKEEJTLUNSEY-UTTGQTJXNA-N |
|---|---|
| Smiles | O=C(C=C1N(CCC2=CC=C(O)C=C2)C(CCC)C(C(=O)OC)C1O)CCCCCCCCCCCCCCCC |
| Superclass | Benzenoids |
| Class | Benzene and substituted derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q8NER1 | TRPV1 | Vanilloid receptor | T83193 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SwissTargetPrediction and SEA |
| P10827 | THRA | Thyroid hormone receptor alpha | T79591 | SEA |
| P10828 | THRB | Thyroid hormone receptor beta-1 | T98933 | SEA |
| P43116 | PTGER2 | Prostanoid EP2 receptor | T38529 | SEA |
| P49841 | GSK3B | Glycogen synthase kinase-3 beta | T70977 | SEA |
| P34972 | CNR2 | Cannabinoid receptor 2 | T37693 | SEA |
| P35408 | PTGER4 | Prostanoid EP4 receptor | T18876 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T83193 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q8NER1 | TRPV1 |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T79591 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10827 | THRA |
| T79591 | DI0197 | Hypo-thyroidism | [ICD-11: 5A00] | P10827 | THRA |
| T98933 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10828 | THRB |
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
| T70977 | DI0288 | Myotonic disorder | [ICD-11: 8C71] | P49841 | GSK3B |
| T37693 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | P34972 | CNR2 |
| T37693 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P34972 | CNR2 |
| T18876 | DI0037 | Asthma | [ICD-11: CA23] | P35408 | PTGER4 |
| T18876 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P35408 | PTGER4 |
| T18876 | DI0264 | Migraine | [ICD-11: 8A80] | P35408 | PTGER4 |
| T18876 | DI0303 | Non-small-cell lung cancer | [ICD-11: 2C25] | P35408 | PTGER4 |
| T18876 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35408 | PTGER4 |
| T18876 | DI0360 | Rectum cancer | [ICD-11: 2B92] | P35408 | PTGER4 |
| T18876 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35408 | PTGER4 |
| T18876 | DI0414 | Transplanted organ/tissue | [ICD-11: QB63] | P35408 | PTGER4 |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
