Compound details
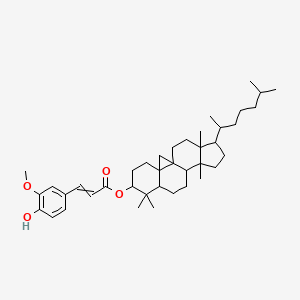
Cycloartanyl ferulate
| Compound ID | CDAMM01156 |
|---|---|
| Common name | Cycloartanyl ferulate | IUPAC name | [7,7,12,16-tetramethyl-15-(6-methylheptan-2-yl)-6-pentacyclo[9.7.0.01,3.03,8.012,16]octadecanyl] 3-(4-hydroxy-3-methoxyphenyl)prop-2-enoate |
| Molecular formula | C40H60O4 |
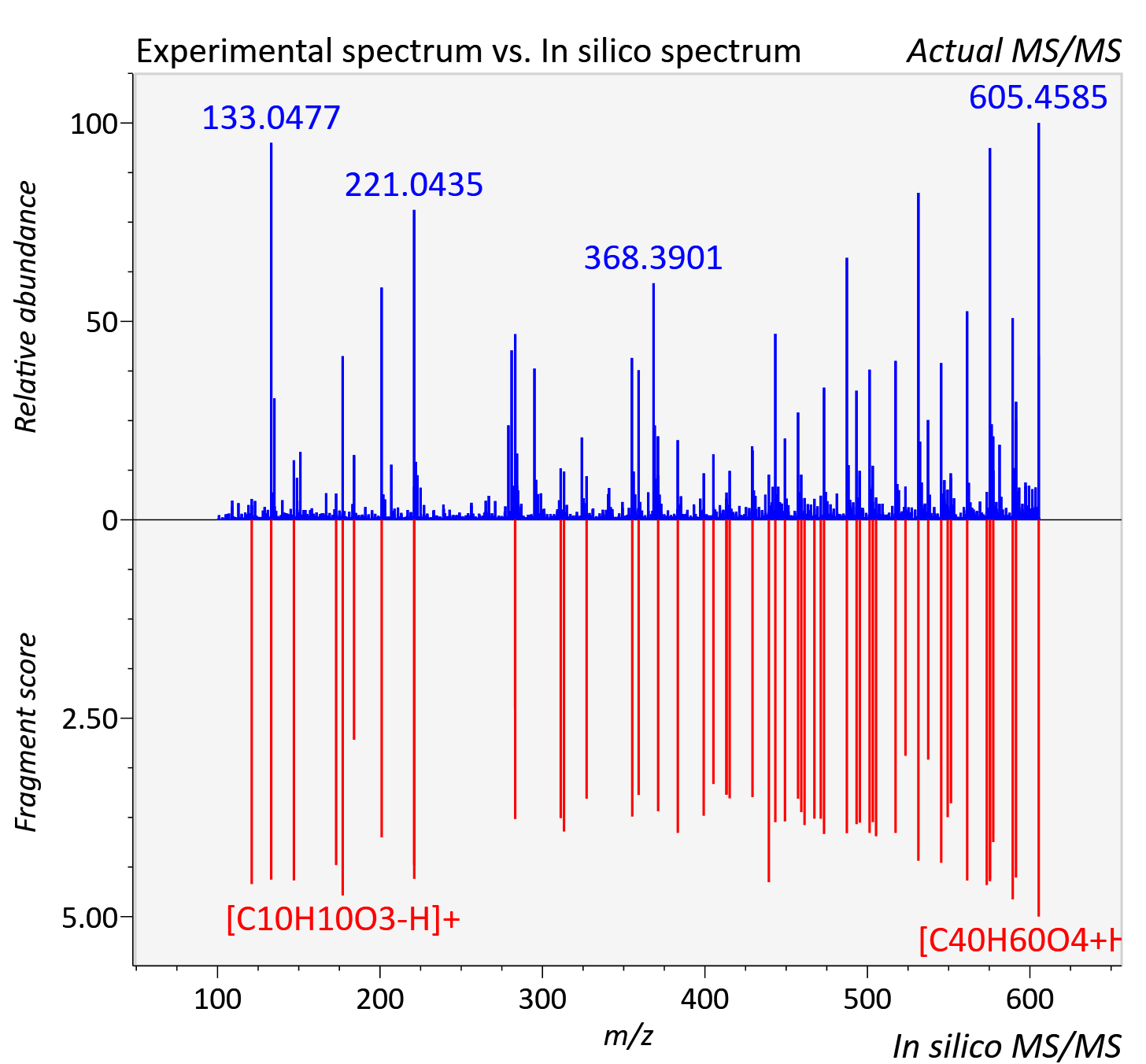
Experimental data
| Retention time | 24.8 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 605.455 | Theoretical mz | 605.456 |
| Error | 1.45 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7132 |
Identifiers and class information
| Inchi key | MFIVAYYVKCBONU-GHRIWEEINA-N |
|---|---|
| Smiles | O=C(OC1CCC23CC43CCC5(C)C(CCC5(C)C4CCC2C1(C)C)C(C)CCCC(C)C)C=CC6=CC=C(O)C(OC)=C6 |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P05067 | APP | Beta amyloid A4 protein | T87024 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| P11388 | TOP2A | DNA topoisomerase II alpha | T17048 | SEA |
| Q09472 | EP300 | Histone acetyltransferase p300 | T25956 | SEA |
| P49810 | PSEN2 | Gamma-secretase | T99204 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P10636 | MAPT | Microtubule-associated protein tau | T45593 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| Q16236 | NFE2L2 | Nuclear factor erythroid 2-related factor 2 | T88505 | SEA |
| P49768 | PSEN1 | Presenilin 1 | T93105 | SEA |
| Q96BI3 | APH1A | Gamma-secretase | T99840 | SEA |
| Q96BI3 | APH1A | Gamma-secretase | T99840 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| Q9P0J0 | NDUFA13 | Mitochondrial complex I | T82391 | SEA |
| Q9Y6M9 | NDUFB9 | HUMAN NADH:ubiquinone oxidoreductase subunit B9 | T01465 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| P03897 | MT-ND3 | NADH dehydrogenase | T77195 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T87024 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P05067 | APP |
| T87024 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P05067 | APP |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T17048 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P11388 | TOP2A |
| T17048 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P11388 | TOP2A |
| T17048 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P11388 | TOP2A |
| T25956 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q09472 | EP300 |
| T25956 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q09472 | EP300 |
| T99204 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P49810 | PSEN2 |
| T99204 | DI0380 | Shoulder lesion | [ICD-11: FB53] | P49810 | PSEN2 |
| T99204 | DI0390 | Soft tissue disorder | [ICD-11: FB56] | P49810 | PSEN2 |
| T45593 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P10636 | MAPT |
| T45593 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P10636 | MAPT |
| T45593 | DI0265 | Mild neurocognitive disorder | [ICD-11: 6D71] | P10636 | MAPT |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
| T88505 | DI0038 | Ataxic disorder | [ICD-11: 8A03] | Q16236 | NFE2L2 |
| T93105 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P49768 | PSEN1 |
| T93105 | DI0380 | Shoulder lesion | [ICD-11: FB53] | P49768 | PSEN1 |
| T93105 | DI0390 | Soft tissue disorder | [ICD-11: FB56] | P49768 | PSEN1 |
| T99840 | DI0119 | Desmoid tumour | [ICD-11: 2F7C] | Q96BI3 | APH1A |
| T99840 | DI0119 | Desmoid tumour | [ICD-11: 2F7C] | Q96BI3 | APH1A |
| T82391 | DI0069 | Cardiomyopathy | [ICD-11: BC43] | Q9P0J0 | NDUFA13 |
| T77195 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P03897 | MT-ND3 |
