Compound details
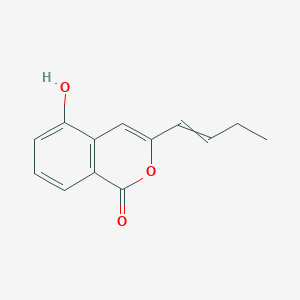
Artemidinol
| Compound ID | CDAMM01154 |
|---|---|
| Common name | Artemidinol | IUPAC name | 3-but-1-enyl-5-hydroxyisochromen-1-one |
| Molecular formula | C13H12O3 |
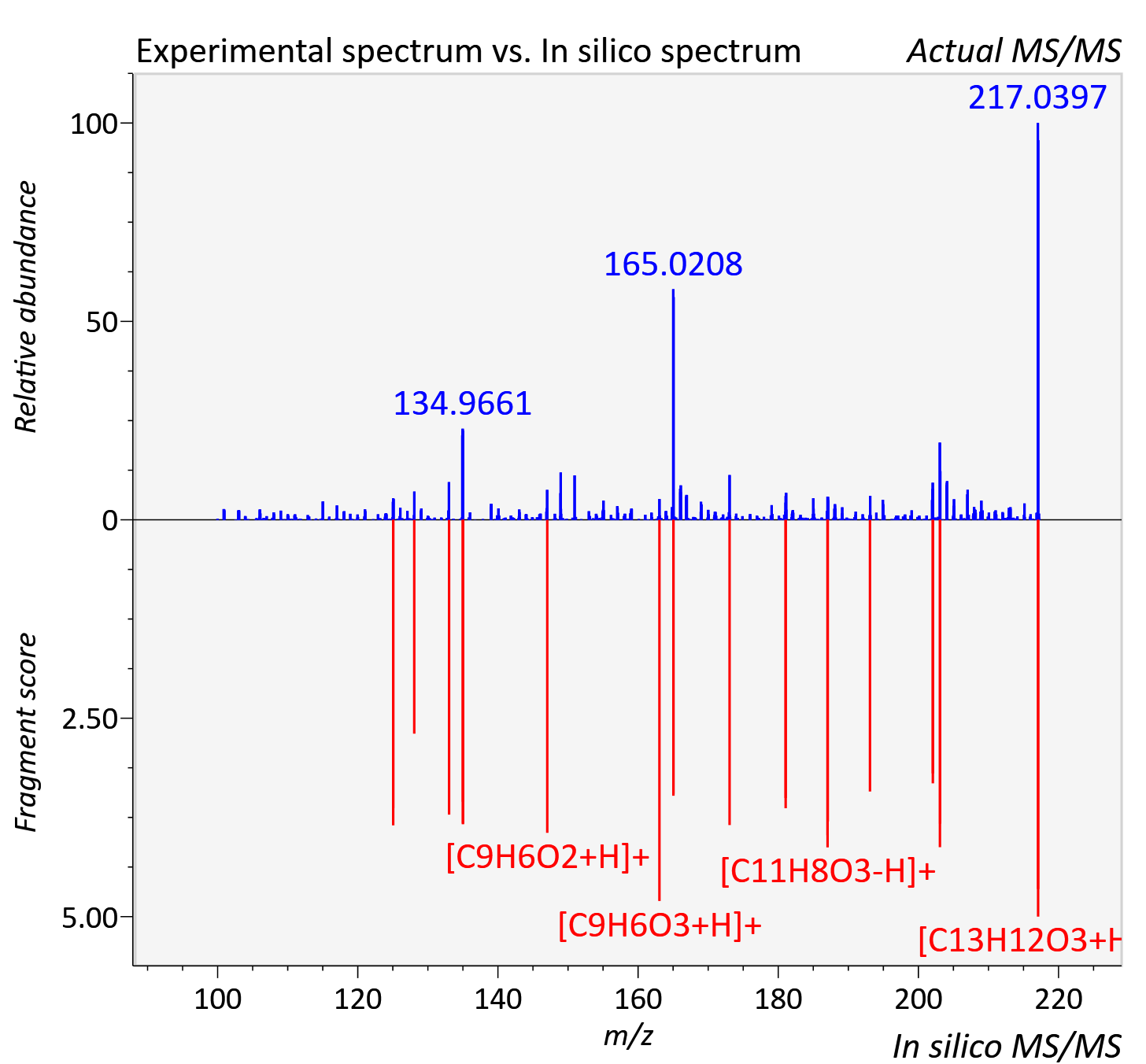
Experimental data
| Retention time | 20.5 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 217.083 | Theoretical mz | 217.086 |
| Error | 14.63 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.9325 |
Identifiers and class information
| Inchi key | QBZHMYUXUVZDQT-HYXAFXHYSA-N |
|---|---|
| Smiles | O=C1OC(C=CCC)=CC=2C(O)=CC=CC12 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Isocoumarins and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| Q16549 | PCSK7 | Subtilisin/kexin type 7 | T33211 | SEA |
| Q86U86 | PBRM1 | Protein polybromo-1 | T00973 | SEA |
| O35273 | SMAD3 | Mothers against decapentaplegic homolog 3 | T35445 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
| H3BPQ1 | TCF4 | Transcription factor 7-like 2 | T54646 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T35445 | DI0226 | Kidney fibrosis | [ICD-11: GC01] | O35273 | SMAD3 |
