Compound details
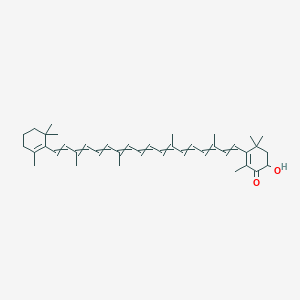
3-Hydroxyechinenone
| Compound ID | CDAMM01123 |
|---|---|
| Common name | 3-Hydroxyechinenone | IUPAC name | 6-hydroxy-2,4,4-trimethyl-3-[3,7,12,16-tetramethyl-18-(2,6,6-trimethylcyclohexen-1-yl)octadeca-1,3,5,7,9,11,13,15,17-nonaenyl]cyclohex-2-en-1-one |
| Molecular formula | C40H54O2 |
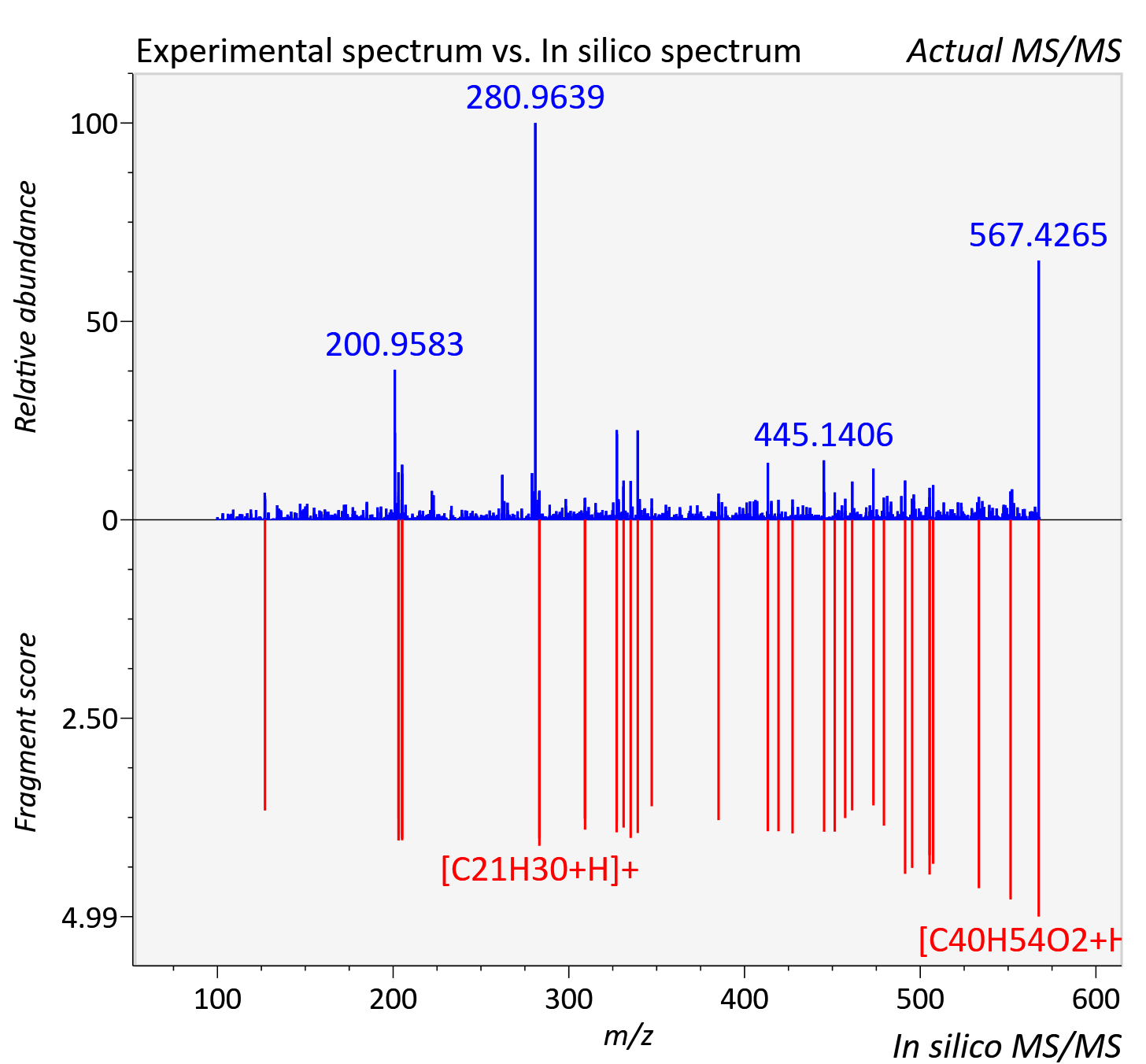
Experimental data
| Retention time | 21.92 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 567.427 | Theoretical mz | 567.419 |
| Error | 14.32 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.8948 |
Identifiers and class information
| Inchi key | DFNMSBYEEKBETA-GVVOHZSFNA-N |
|---|---|
| Smiles | O=C1C(=C(C=CC(=CC=CC(=CC=CC=C(C=CC=C(C=CC2=C(C)CCCC2(C)C)C)C)C)C)C(C)(C)CC1O)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P10276 | RARA | Retinoic acid receptor alpha | T94085 | SEA |
| O43174 | CYP26A1 | Cytochrome P450 26A1 | T84634 | SEA |
| P19793 | RXRA | Retinoid X receptor alpha (by homology) | T13726 | SEA |
| P02753 | RBP4 | Plasma retinol-binding protein | T55703 | SEA |
| P35398 | RORA | Nuclear receptor ROR-alpha | T43206 | SEA |
| P28702 | RXRB | Retinoid X receptor beta | T60077 | SEA |
| P13631 | RARG | Retinoic acid receptor gamma | T82146 | SEA |
| P48443 | RXRG | Retinoid X receptor gamma | T46456 | SEA |
| P10826 | RARB | Retinoic acid receptor beta | T61657 | SEA |
| Q92753 | RORB | Nuclear receptor ROR-beta | T48812 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T94085 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P10276 | RARA |
| T94085 | DI0251 | Mature T-cell lymphoma | [ICD-11: 2A90] | P10276 | RARA |
| T84634 | DI0314 | Oesophagitis | [ICD-11: DA24] | O43174 | CYP26A1 |
| T13726 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P19793 | RXRA |
| T13726 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P19793 | RXRA |
| T55703 | DI0212 | Inherited retinal dystrophy | [ICD-11: 9B70] | P02753 | RBP4 |
| T43206 | DI0235 | Liver cancer | [ICD-11: 2C12] | P35398 | RORA |
| T43206 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P35398 | RORA |
| T60077 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P28702 | RXRB |
| T82146 | DI0005 | Acne vulgaris | [ICD-11: ED80] | P13631 | RARG |
| T82146 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P13631 | RARG |
| T82146 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P13631 | RARG |
| T82146 | DI0277 | Muscle calcification/ossification | [ICD-11: FB31] | P13631 | RARG |
| T82146 | DI0351 | Psoriasis | [ICD-11: EA90] | P13631 | RARG |
| T46456 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P48443 | RXRG |
| T46456 | DI0232 | Light sensitivity impairment | [ICD-11: 9D45] | P48443 | RXRG |
| T61657 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P10826 | RARB |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
