Compound details
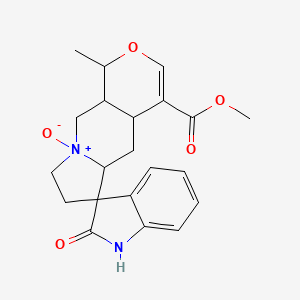
Isomitraphylline N-oxide
| Compound ID | CDAMM01116 |
|---|---|
| Common name | Isomitraphylline N-oxide | IUPAC name | methyl 1-methyl-9-oxido-2\'-oxospiro[1,4a,5,5a,7,8,10,10a-octahydropyrano[3,4-f]indolizin-9-ium-6,3\'-1H-indole]-4-carboxylate |
| Molecular formula | C21H24N2O5 |
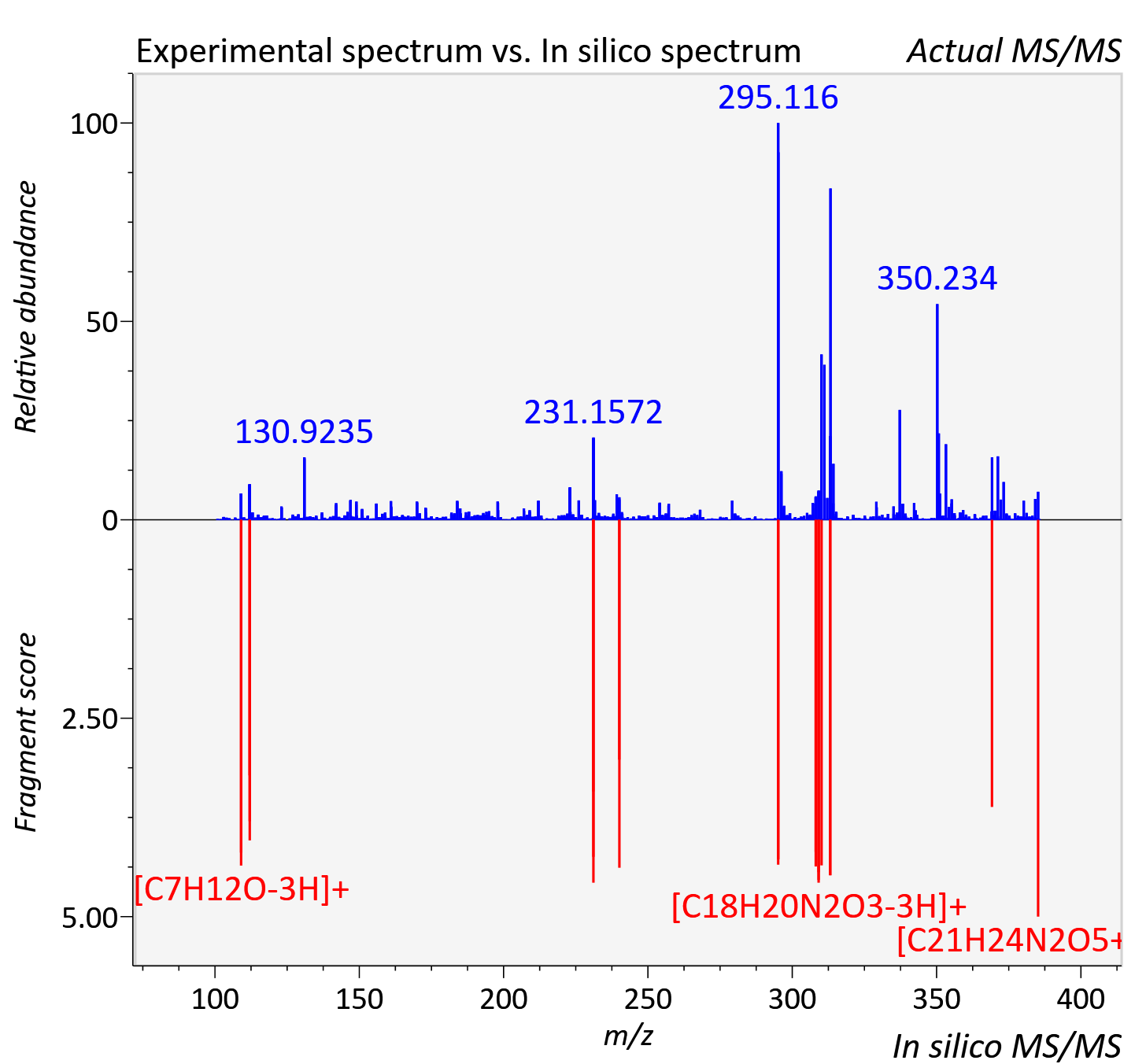
Experimental data
| Retention time | 12.25 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 385.177 | Theoretical mz | 385.176 |
| Error | 2.92 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5669 |
Identifiers and class information
| Inchi key | DTEXPPFMGPLSPX-CKQZDEODNA-N |
|---|---|
| Smiles | O=C(OC)C1=COC(C)C2CN3(=O)CCC4(C(=O)NC=5C=CC=CC54)C3CC12 |
| Superclass | Organoheterocyclic compounds |
| Class | Indolizidines |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T59328 | DI0030 | Angina pectoris | [ICD-11: BA40] | P00533 | EGFR |
| T59328 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P00533 | EGFR |
| T59328 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P00533 | EGFR |
| T59328 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P00533 | EGFR |
| T59328 | DI0220 | Ischemia | [ICD-11: 8B10-8B11] | P00533 | EGFR |
| T59328 | DI0238 | Lung cancer | [ICD-11: 2C25] | P00533 | EGFR |
| T59328 | DI0303 | Non-small-cell lung cancer | [ICD-11: 2C25] | P00533 | EGFR |
| T59328 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P00533 | EGFR |
| T59328 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P00533 | EGFR |
| T59328 | DI0420 | Unspecific body region injury | [ICD-11: ND56] | P00533 | EGFR |
| T75498 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P16083 | NQO2 |
