Compound details
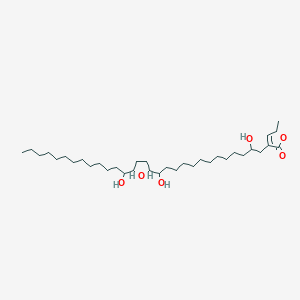
Murisolin
| Compound ID | CDAMM01102 |
|---|---|
| Common name | Murisolin | IUPAC name | 4-[2,13-dihydroxy-13-[5-(1-hydroxytridecyl)oxolan-2-yl]tridecyl]-2-methyl-2H-furan-5-one |
| Molecular formula | C35H64O6 |
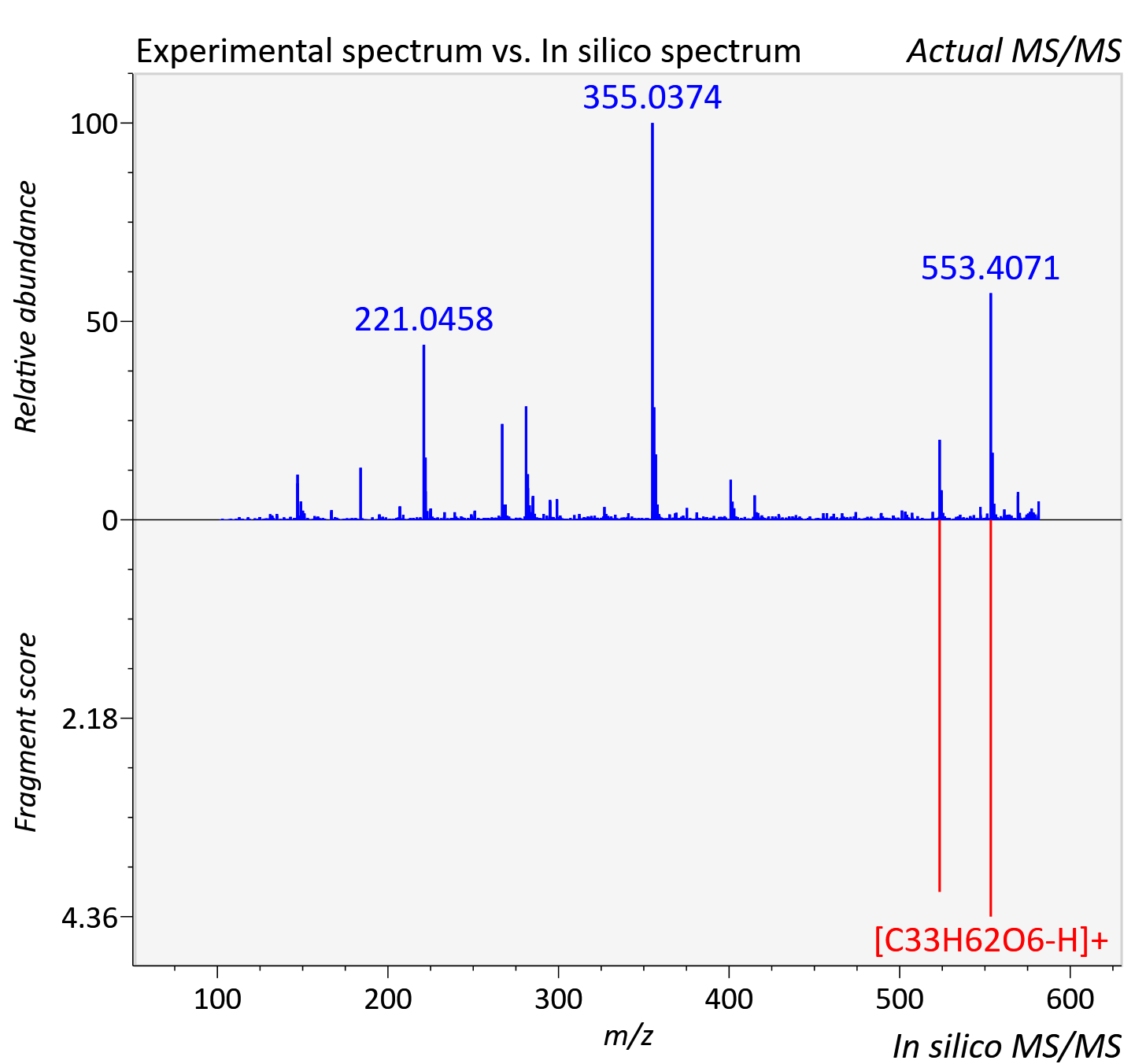
Experimental data
| Retention time | 19.38 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 581.477 | Theoretical mz | 581.477 |
| Error | 0.65 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.4381 |
Identifiers and class information
| Inchi key | PYNFAPLXMQHUNR-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(C=C1CC(O)CCCCCCCCCCC(O)C2OC(CC2)C(O)CCCCCCCCCCCC)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
