Compound details
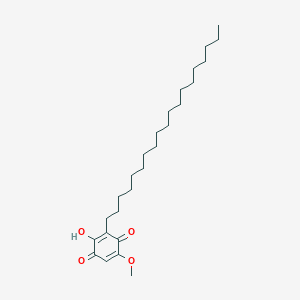
Irisoquin C
| Compound ID | CDAMM01055 |
|---|---|
| Common name | Irisoquin C | IUPAC name | 2-hydroxy-5-methoxy-3-nonadecylcyclohexa-2,5-diene-1,4-dione |
| Molecular formula | C26H44O4 |
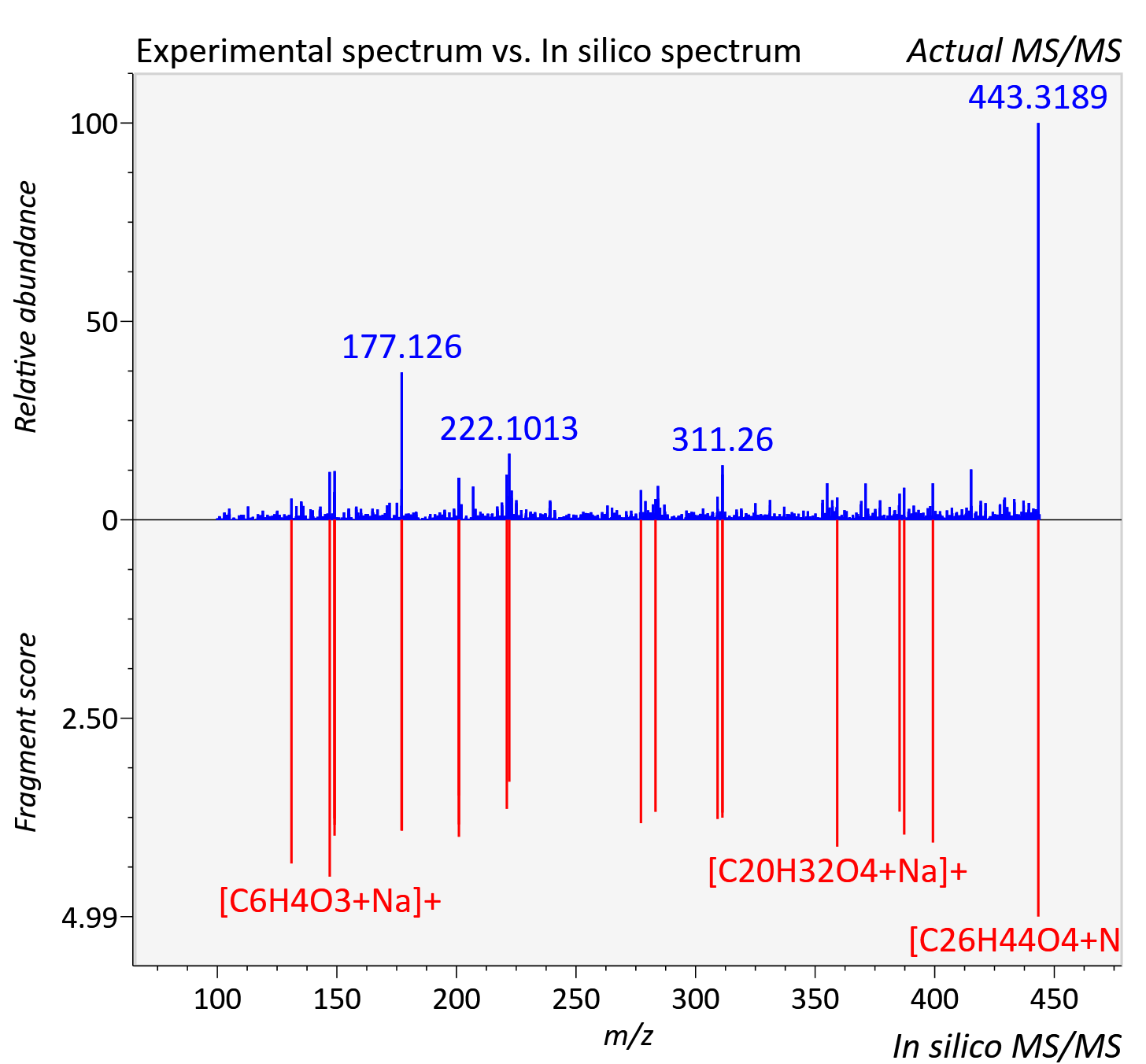
Experimental data
| Retention time | 17.92 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 443.318 | Theoretical mz | 443.313 |
| Error | 11.24 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5289 |
Identifiers and class information
| Inchi key | SUAQMIMGKZVLBE-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C1C=C(OC)C(=O)C(=C1O)CCCCCCCCCCCCCCCCCCC |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P10635 | CYP2D6 | Cytochrome P450 2D6 | T57392 | SwissTargetPrediction |
| P11712 | CYP2C9 | Cytochrome P450 2C9 | T19244 | SwissTargetPrediction |
| P08684 | CYP3A4 | Cytochrome P450 3A4 | T37848 | SwissTargetPrediction |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SwissTargetPrediction |
| P98170 | XIAP | Inhibitor of apoptosis protein 3 | T16769 | SwissTargetPrediction |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SwissTargetPrediction |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SwissTargetPrediction and SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SwissTargetPrediction |
| P10827 | THRA | Thyroid hormone receptor alpha | T79591 | SEA |
| P10828 | THRB | Thyroid hormone receptor beta-1 | T98933 | SEA |
| P27695 | APEX1 | DNA-(apurinic or apyrimidinic site) lyase | T13348 | SEA |
| Q92831 | KAT2B | Histone acetyltransferase PCAF | T16902 | SwissTargetPrediction |
| P08842 | STS | Steryl-sulfatase | T33489 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P10826 | RARB | Retinoic acid receptor beta | T61657 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| Q9NQS5 | GPR84 | G-protein coupled receptor 84 | T98091 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T57392 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P10635 | CYP2D6 |
| T19244 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P11712 | CYP2C9 |
| T37848 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P08684 | CYP3A4 |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T16769 | DI0259 | Metastatic lymph node neoplasm | [ICD-11: 2D60] | P98170 | XIAP |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T79591 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10827 | THRA |
| T79591 | DI0197 | Hypo-thyroidism | [ICD-11: 5A00] | P10827 | THRA |
| T98933 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10828 | THRB |
| T13348 | DI0365 | Retinopathy | [ICD-11: 9B71] | P27695 | APEX1 |
| T13348 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P27695 | APEX1 |
| T33489 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P08842 | STS |
| T61657 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P10826 | RARB |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T98091 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q9NQS5 | GPR84 |
| T98091 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q9NQS5 | GPR84 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
