Compound details
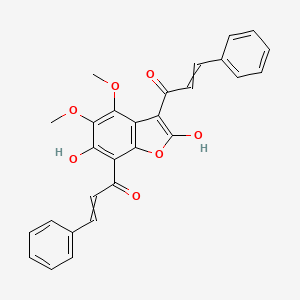
Didymocalyxin B
| Compound ID | CDAMM01049 |
|---|---|
| Common name | Didymocalyxin B | IUPAC name | 1-[2,6-dihydroxy-4,5-dimethoxy-7-(3-phenylprop-2-enoyl)-1-benzofuran-3-yl]-3-phenylprop-2-en-1-one |
| Molecular formula | C28H22O7 |
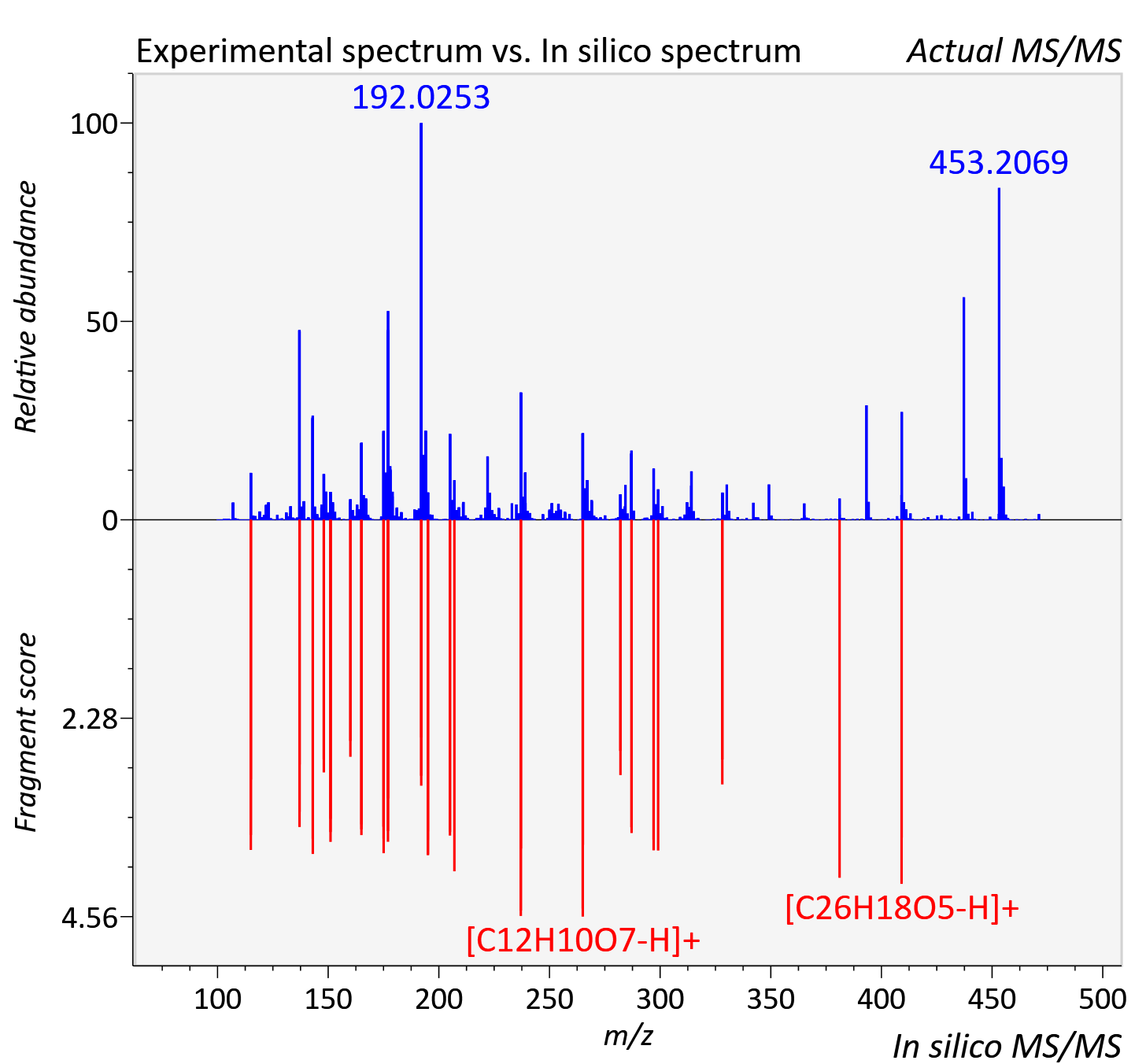
Experimental data
| Retention time | 0.29 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 471.146 | Theoretical mz | 471.144 |
| Error | 3.02 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.957 |
Identifiers and class information
| Inchi key | SYJASFPLLHXQRX-VZZMMMTHSA-N |
|---|---|
| Smiles | O=C1OC=2C(C(=O)C=CC=3C=CC=CC3)=C(O)C(OC)=C(OC)C2C1=C(O)C=CC=4C=CC=CC4 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Linear 1,3-diarylpropanoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P20839 | IMPDH1 | Inosine-5'-monophosphate dehydrogenase 1 | T40111 | SEA |
| P12268 | IMPDH2 | Inosine-5'-monophosphate dehydrogenase 2 | T89360 | SEA |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| P22001 | KCNA3 | Voltage-gated potassium channel subunit Kv1.3 | T76914 | SwissTargetPrediction and SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P19438 | TNFRSF1A | Tumor necrosis factor receptor R1 | T86552 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| O75884 | RBBP9 | Putative hydrolase RBBP9 | T12084 | SEA |
| Q9H4B7 | TUBB1 | Tubulin beta-1 chain | T84397 | SEA |
| O43826 | SLC37A4 | Glucose-6-phosphate translocase | T47306 | SEA |
| P16220 | CREB1 | Cyclic AMP-responsive element-binding protein | T92098 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T40111 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P20839 | IMPDH1 |
| T40111 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P20839 | IMPDH1 |
| T40111 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | P20839 | IMPDH1 |
| T40111 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P20839 | IMPDH1 |
| T40111 | DI0413 | Transplant rejection | [ICD-11: NE84] | P20839 | IMPDH1 |
| T89360 | DI0413 | Transplant rejection | [ICD-11: NE84] | P12268 | IMPDH2 |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T76914 | DI0198 | Idiopathic inflammatory myopathy | [ICD-11: 4A41] | P22001 | KCNA3 |
| T76914 | DI0351 | Psoriasis | [ICD-11: EA90] | P22001 | KCNA3 |
| T76914 | DI0352 | Psoriatic arthritis | [ICD-11: FA21] | P22001 | KCNA3 |
| T86552 | DI0060 | Brain cancer | [ICD-11: 2A00] | P19438 | TNFRSF1A |
| T86552 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P19438 | TNFRSF1A |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T84397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9H4B7 | TUBB1 |
