Compound details
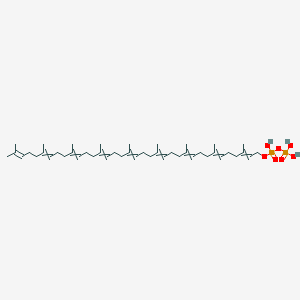
Solanesyl-PP
| Compound ID | CDAMM01018 |
|---|---|
| Common name | Solanesyl-PP | IUPAC name | 3,7,11,15,19,23,27,31,35-nonamethylhexatriaconta-2,6,10,14,18,22,26,30,34-nonaenyl phosphono hydrogen phosphate |
| Molecular formula | C45H76O7P2 |
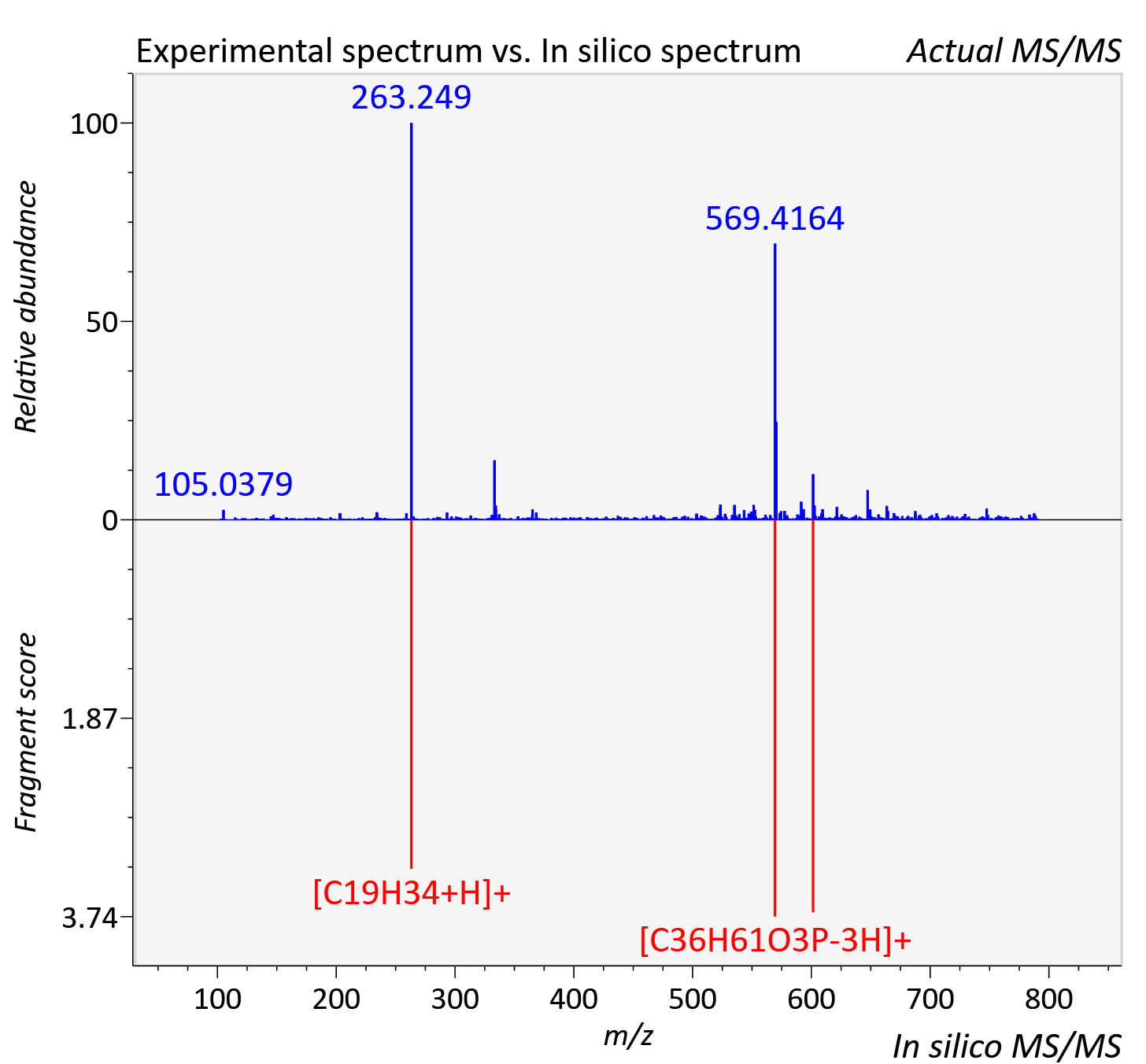
Experimental data
| Retention time | 15.87 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 791.523 | Theoretical mz | 791.514 |
| Error | 10.77 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.8306 |
Identifiers and class information
| Inchi key | IVLBHBFTRNVIAP-HUIBRQQWNA-N |
|---|---|
| Smiles | O=P(O)(O)OP(=O)(O)OCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)CCC=C(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P48449 | LSS | Lanosterol synthase | T56175 | SEA |
| Q14534 | SQLE | Squalene monooxygenase (by homology) | T93344 | SEA |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SwissTargetPrediction and SEA |
| P53609 | PGGT1B | Geranylgeranyl transferase type I | T76396 | SwissTargetPrediction and SEA |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P53602 | MVD | Diphosphomevalonate decarboxylase | T96862 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T56175 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | P48449 | LSS |
| T93344 | DI0118 | Dermatophytosis | [ICD-11: 1F28] | Q14534 | SQLE |
| T93344 | DI0385 | Skin fungal infection disorder | [ICD-11: EA60] | Q14534 | SQLE |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T76396 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P53609 | PGGT1B |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
