Compound details
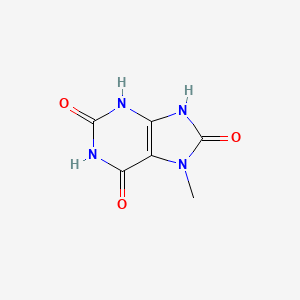
7-Methyluric acid
| Compound ID | CDAMM00992 |
|---|---|
| Common name | 7-Methyluric acid | IUPAC name | 7-methyl-3,9-dihydropurine-2,6,8-trione |
| Molecular formula | C6H6N4O3 |
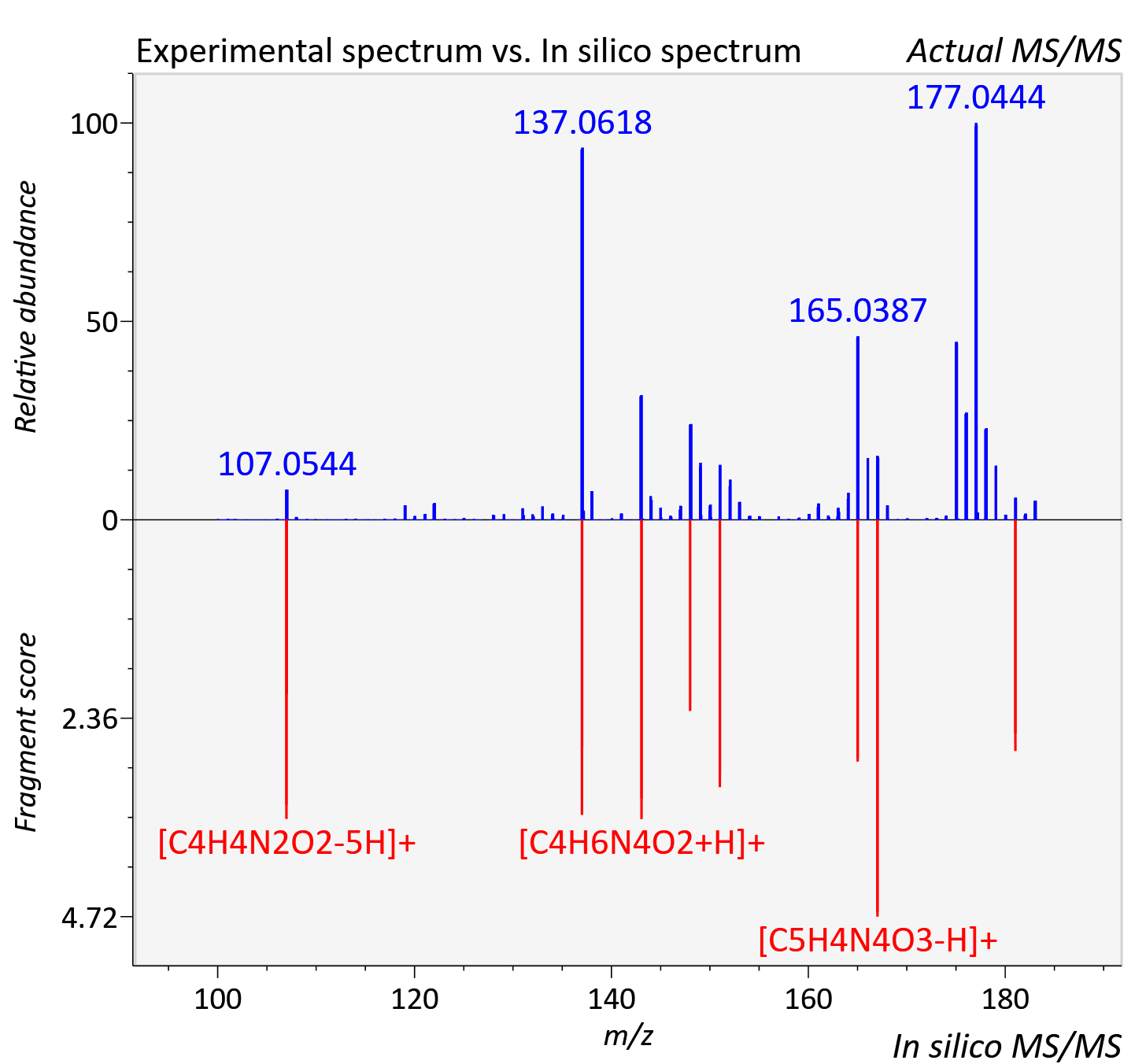
Experimental data
| Retention time | 0.29 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 183.049 | Theoretical mz | 183.051 |
| Error | 10.25 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.8127 |
Identifiers and class information
| Inchi key | YHNNPKUFPWLTOP-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C1NC(=O)C2=C(N1)NC(=O)N2C |
| Superclass | Organoheterocyclic compounds |
| Class | Imidazopyrimidines |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P22303 | ACHE | Acetylcholinesterase | T30082 | SwissTargetPrediction |
| Q12809 | KCNH2 | HERG | T20251 | SwissTargetPrediction |
| P0DMS8 | ADORA3 | Adenosine A3 receptor | T36059 | SwissTargetPrediction |
| Q9Y2T3 | GDA | Guanine deaminase | T92460 | SwissTargetPrediction and SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T30082 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P22303 | ACHE |
| T30082 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22303 | ACHE |
| T30082 | DI0282 | Myasthenia gravis | [ICD-11: 8C6Y] | P22303 | ACHE |
| T30082 | DI0313 | Oesophageal/gastroduodenal disorder | [ICD-11: DD90] | P22303 | ACHE |
| T30082 | DI0332 | Pediculosis | [ICD-11: 1G00] | P22303 | ACHE |
| T30082 | DI0421 | Unspecific substance harmful effect | [ICD-11: NE6Z] | P22303 | ACHE |
| T20251 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | Q12809 | KCNH2 |
| T36059 | DI0351 | Psoriasis | [ICD-11: EA90] | P0DMS8 | ADORA3 |
| T36059 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P0DMS8 | ADORA3 |
