Compound details
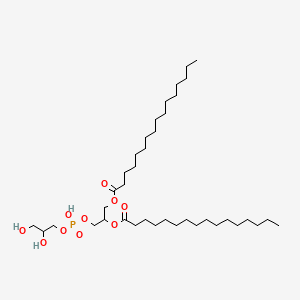
PG(16:0/16:0)
| Compound ID | CDAMM00989 |
|---|---|
| Common name | PG(16:0/16:0) | IUPAC name | [3-[2,3-dihydroxypropoxy(hydroxy)phosphoryl]oxy-2-hexadecanoyloxypropyl] hexadecanoate |
| Molecular formula | C38H75O10P |
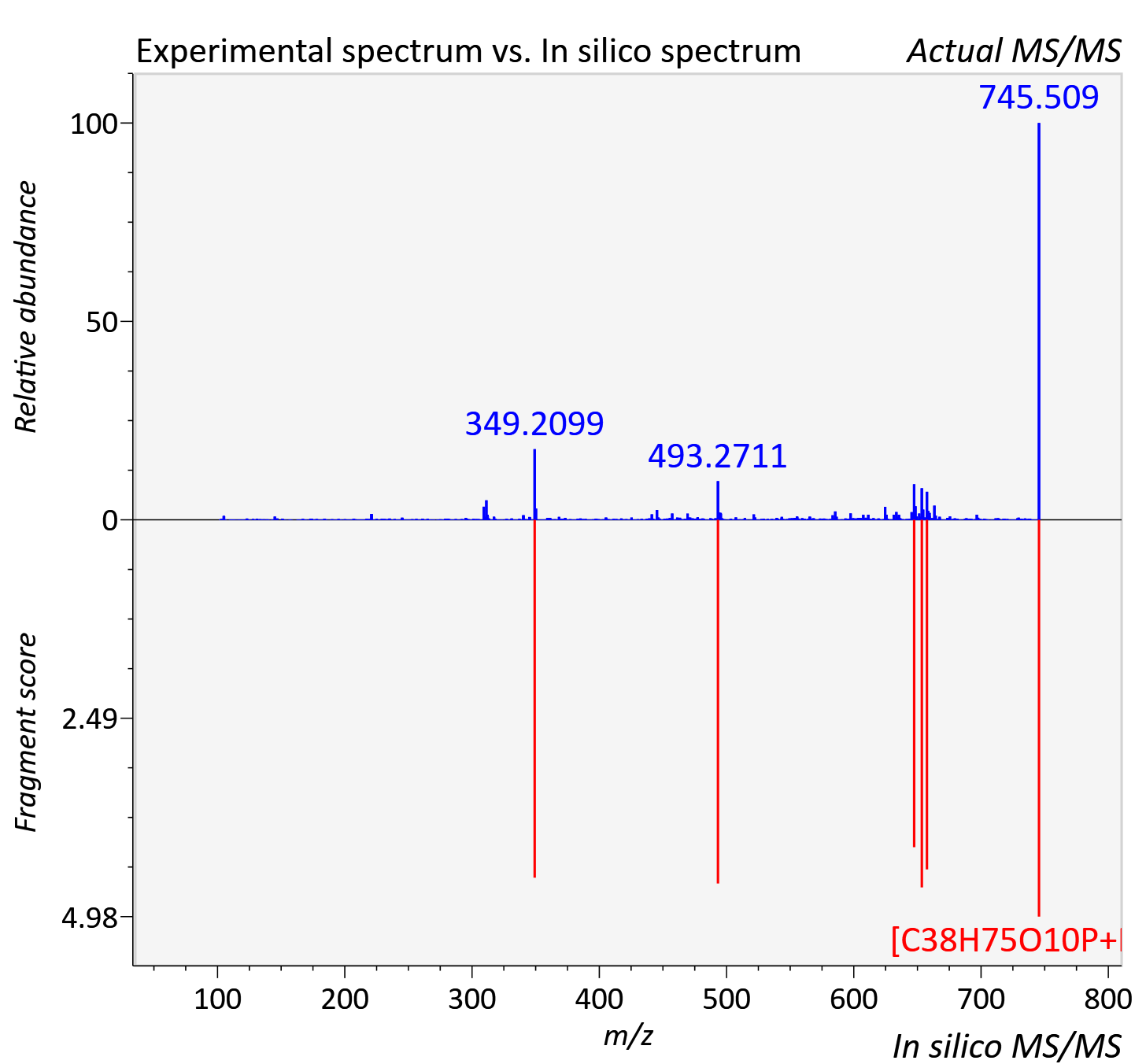
Experimental data
| Retention time | 15.61 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 745.508 | Theoretical mz | 745.499 |
| Error | 12.18 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.9915 |
Identifiers and class information
| Inchi key | BIABMEZBCHDPBV-MPQUPPDSSA-N |
|---|---|
| Smiles | O=C(OCC(OC(=O)CCCCCCCCCCCCCCC)COP(=O)(O)OCC(O)CO)CCCCCCCCCCCCCCC |
| Superclass | Lipids and lipid-like molecules |
| Class | Glycerophospholipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| O00206 | TLR4 | Toll-like receptor 4 (by homology) | T81443 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P16109 | SELP | P-selectin | T10965 | SEA |
| Q9H228 | S1PR5 | Sphingosine 1-phosphate receptor Edg-8 | T50089 | SEA |
| P35813 | PPM1A | Protein phosphatase 2C alpha | T55260 | SEA |
| Q13822 | ENPP2 | Autotaxin | T63512 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| Q99500 | S1PR3 | Sphingosine 1-phosphate receptor Edg-3 | T11241 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9H1C0 | LPAR5 | Lysophosphatidic acid receptor 5 | T18616 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| O95977 | S1PR4 | Sphingosine 1-phosphate receptor Edg-6 | T17523 | SEA |
| O60906 | SMPD2 | Neutral sphingomyelinase | T57316 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T81443 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | O00206 | TLR4 |
| T81443 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O00206 | TLR4 |
| T81443 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | O00206 | TLR4 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T10965 | DI0088 | Circulatory system disease | [ICD-11: BE2Z] | P16109 | SELP |
| T10965 | DI0381 | Sickle-cell disorder | [ICD-11: 3A51] | P16109 | SELP |
| T50089 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | Q9H228 | S1PR5 |
| T55260 | DI0268 | Molluscum contagiosum | [ICD-11: 1E76] | P35813 | PPM1A |
| T63512 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q13822 | ENPP2 |
| T11241 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q99500 | S1PR3 |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
