Compound details
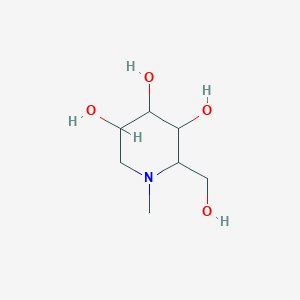
N-Methyl-1-deoxynojirimycin
| Compound ID | CDAMM00977 |
|---|---|
| Common name | N-Methyl-1-deoxynojirimycin | IUPAC name | 2-(hydroxymethyl)-1-methylpiperidine-3,4,5-triol |
| Molecular formula | C7H15NO4 |
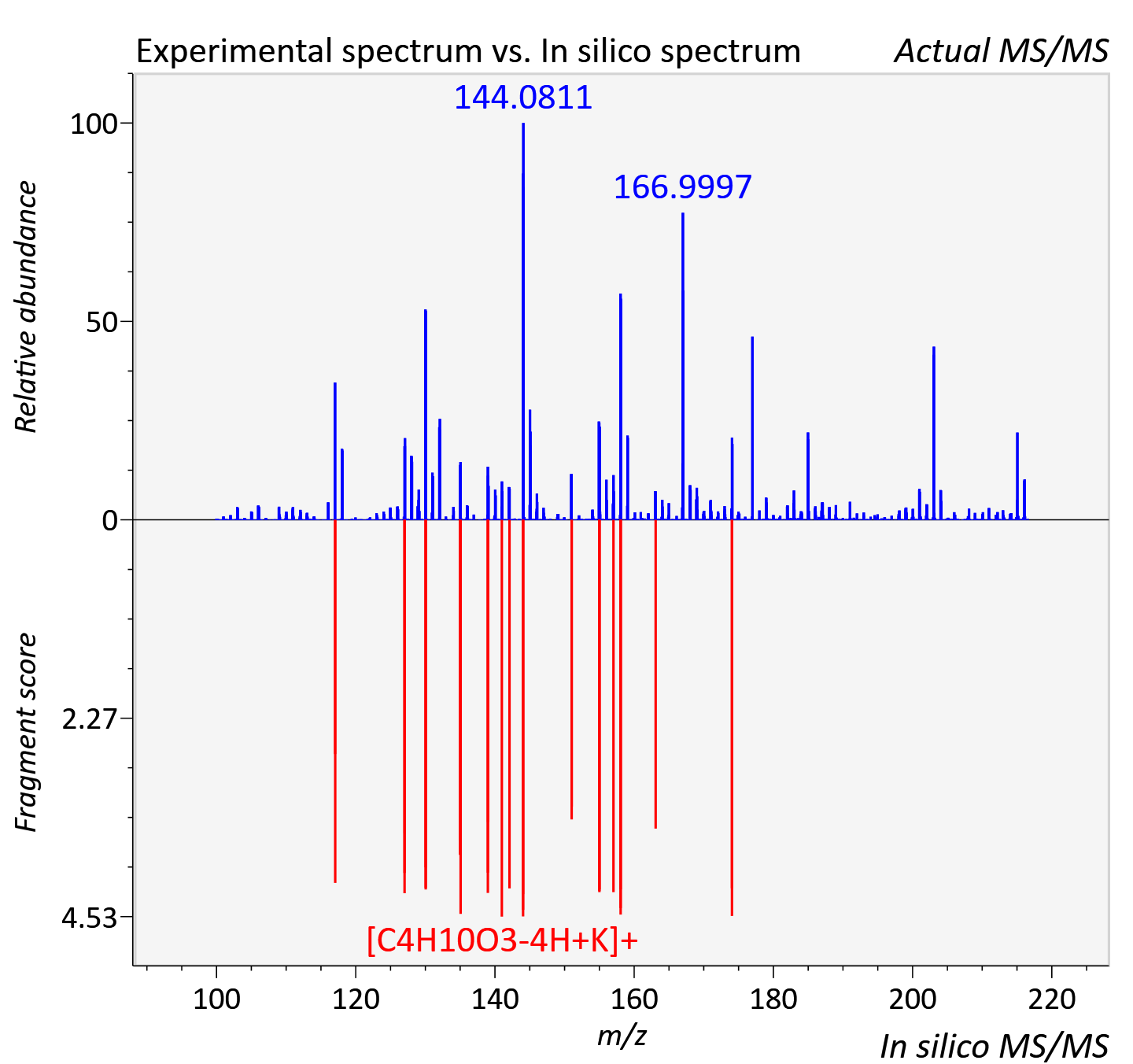
Experimental data
| Retention time | 0.43 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 216.062 | Theoretical mz | 216.063 |
| Error | 3.97 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.6872 |
Identifiers and class information
| Inchi key | AAKDPDFZMNYDLR-UHFFFAOYNA-N |
|---|---|
| Smiles | OCC1N(C)CC(O)C(O)C1O |
| Superclass | Organoheterocyclic compounds |
| Class | Piperidines |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P11940 | PABPC1 | Polyadenylate-binding protein 1 | T58679 | SwissTargetPrediction |
| O00754 | MAN2B1 | Lysosomal alpha-mannosidase | T63156 | SwissTargetPrediction and SEA |
| P06280 | GLA | Alpha-galactosidase A | T61339 | SwissTargetPrediction |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SwissTargetPrediction |
| P04054 | PLA2G1B | Phospholipase A2 group 1B | T31479 | SwissTargetPrediction |
| P48449 | LSS | Lanosterol synthase | T56175 | SwissTargetPrediction |
| P16278 | GLB1 | Beta-galactosidase (by homology) | T73134 | SwissTargetPrediction and SEA |
| Q16739 | UGCG | Ceramide glucosyltransferase | T14908 | SwissTargetPrediction and SEA |
| O43451 | MGAM | Maltase-glucoamylase | T92777 | SwissTargetPrediction |
| P10253 | GAA | Lysosomal alpha-glucosidase (by homology) | T31514 | SwissTargetPrediction and SEA |
| P06280 | GLA | Alpha-galactosidase A | T61339 | SwissTargetPrediction |
| P06280 | GLA | Alpha-galactosidase A | T61339 | SwissTargetPrediction and SEA |
| P04062 | GBA | Beta-glucocerebrosidase | T84173 | SwissTargetPrediction |
| P06737 | PYGL | Liver glycogen phosphorylase | T19184 | SwissTargetPrediction |
| Q9UMJ8 | GBA1 | Lysosomal acid glucosylceramidase | T63243 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T58679 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P11940 | PABPC1 |
| T63156 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | O00754 | MAN2B1 |
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T31479 | DI0230 | Leishmaniasis | [ICD-11: 1F54] | P04054 | PLA2G1B |
| T31479 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P04054 | PLA2G1B |
| T31479 | DI0367 | Rosacea | [ICD-11: ED90] | P04054 | PLA2G1B |
| T31479 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P04054 | PLA2G1B |
| T56175 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | P48449 | LSS |
| T73134 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | P16278 | GLB1 |
| T14908 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q16739 | UGCG |
| T14908 | DI0257 | Metabolic disorder | [ICD-11: 5C50-5D2Z] | Q16739 | UGCG |
| T92777 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | O43451 | MGAM |
| T92777 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | O43451 | MGAM |
| T31514 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P10253 | GAA |
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T84173 | DI0257 | Metabolic disorder | [ICD-11: 5C50-5D2Z] | P04062 | GBA |
| T63243 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q9UMJ8 | GBA1 |
