Compound details
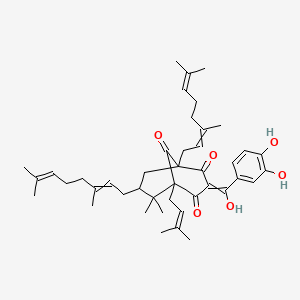
Oblongifolin D;(+)-Oblongifolin D
| Compound ID | CDAMM00976 |
|---|---|
| Common name | Oblongifolin D;(+)-Oblongifolin D | IUPAC name | 3-[(3,4-dihydroxyphenyl)-hydroxymethylidene]-1,7-bis(3,7-dimethylocta-2,6-dienyl)-6,6-dimethyl-5-(3-methylbut-2-enyl)bicyclo[3.3.1]nonane-2,4,9-trione |
| Molecular formula | C43H58O6 |
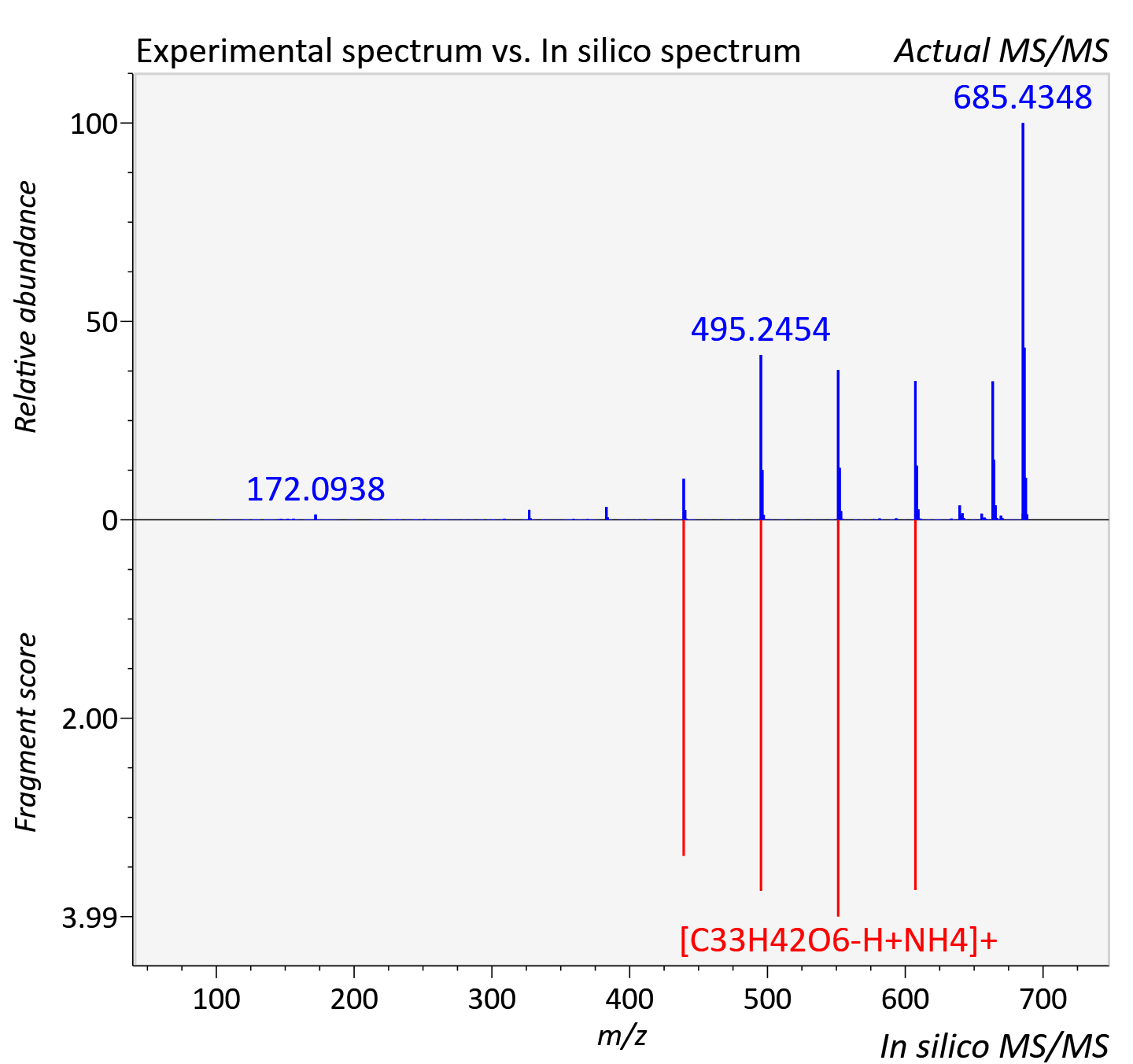
Experimental data
| Retention time | 20.31 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 688.447 | Theoretical mz | 688.457 |
| Error | 13.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7425 |
Identifiers and class information
| Inchi key | OGBLTDAGYQWRIK-XGCSDDDWNA-N |
|---|---|
| Smiles | O=C(C1=CC=C(O)C(O)=C1)C=2C(=O)C3(C(=O)C(C2O)(CC=C(C)CCC=C(C)C)CC(CC=C(C)CCC=C(C)C)C3(C)C)CC=C(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q13133 | NR1H3 | LXR-alpha | T52297 | SwissTargetPrediction |
| P07858 | CTSB | Cathepsin (B and K) | T61746 | SwissTargetPrediction |
| Q09472 | EP300 | Histone acetyltransferase p300 | T25956 | SwissTargetPrediction and SEA |
| Q92831 | KAT2B | Histone acetyltransferase PCAF | T16902 | SwissTargetPrediction |
| P08581 | MET | Hepatocyte growth factor receptor | T40474 | SwissTargetPrediction |
| Q96EB6 | SIRT1 | NAD-dependent deacetylase sirtuin 1 | T14731 | SwissTargetPrediction |
| P42224 | STAT1 | Signal transducer and activator of transcription 1-alpha/beta | T64205 | SwissTargetPrediction |
| P08311 | CTSG | Cathepsin G | T86385 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T52297 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q13133 | NR1H3 |
| T61746 | DI0023 | Alopecia | [ICD-11: ED70] | P07858 | CTSB |
| T61746 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P07858 | CTSB |
| T61746 | DI0037 | Asthma | [ICD-11: CA23] | P07858 | CTSB |
| T61746 | DI0055 | Bone cancer | [ICD-11: 2B5Z] | P07858 | CTSB |
| T61746 | DI0060 | Brain cancer | [ICD-11: 2A00] | P07858 | CTSB |
| T61746 | DI0067 | Cardiac arrest | [ICD-11: MC82] | P07858 | CTSB |
| T61746 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | P07858 | CTSB |
| T61746 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P07858 | CTSB |
| T61746 | DI0087 | Chronic pain | [ICD-11: MG30] | P07858 | CTSB |
| T61746 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P07858 | CTSB |
| T61746 | DI0124 | Digestive system disease | [ICD-11: DE2Z] | P07858 | CTSB |
| T61746 | DI0178 | Hepatic fibrosis/cirrhosis | [ICD-11: DB93] | P07858 | CTSB |
| T61746 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P07858 | CTSB |
| T61746 | DI0253 | Meningioma | [ICD-11: 2A01] | P07858 | CTSB |
| T61746 | DI0262 | Metastatic tumour | [ICD-11: 2D50-2E2Z] | P07858 | CTSB |
| T61746 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P07858 | CTSB |
| T61746 | DI0296 | Neurodegenerative disorder | [ICD-11: 8A20-8A23] | P07858 | CTSB |
| T61746 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P07858 | CTSB |
| T61746 | DI0351 | Psoriasis | [ICD-11: EA90] | P07858 | CTSB |
| T61746 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P07858 | CTSB |
| T25956 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q09472 | EP300 |
| T25956 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q09472 | EP300 |
| T40474 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08581 | MET |
| T40474 | DI0303 | Non-small-cell lung cancer | [ICD-11: 2C25] | P08581 | MET |
| T40474 | DI0407 | Thyroid cancer | [ICD-11: 2D10] | P08581 | MET |
| T14731 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q96EB6 | SIRT1 |
| T14731 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q96EB6 | SIRT1 |
| T14731 | DI0431 | Vasculitis | [ICD-11: 4A44] | Q96EB6 | SIRT1 |
| T64205 | DI0383 | Skin and skin-structure infection | [ICD-11: 1F28-1G0Z] | P42224 | STAT1 |
| T86385 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P08311 | CTSG |
