Compound details
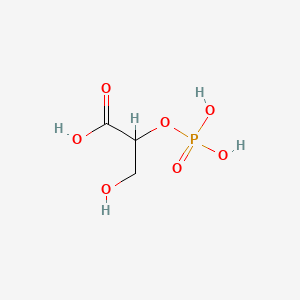
2-Phosphoglyceric acid
| Compound ID | CDAMM00964 |
|---|---|
| Common name | 2-Phosphoglyceric acid | IUPAC name | 3-hydroxy-2-phosphonooxypropanoic acid |
| Molecular formula | C3H7O7P |
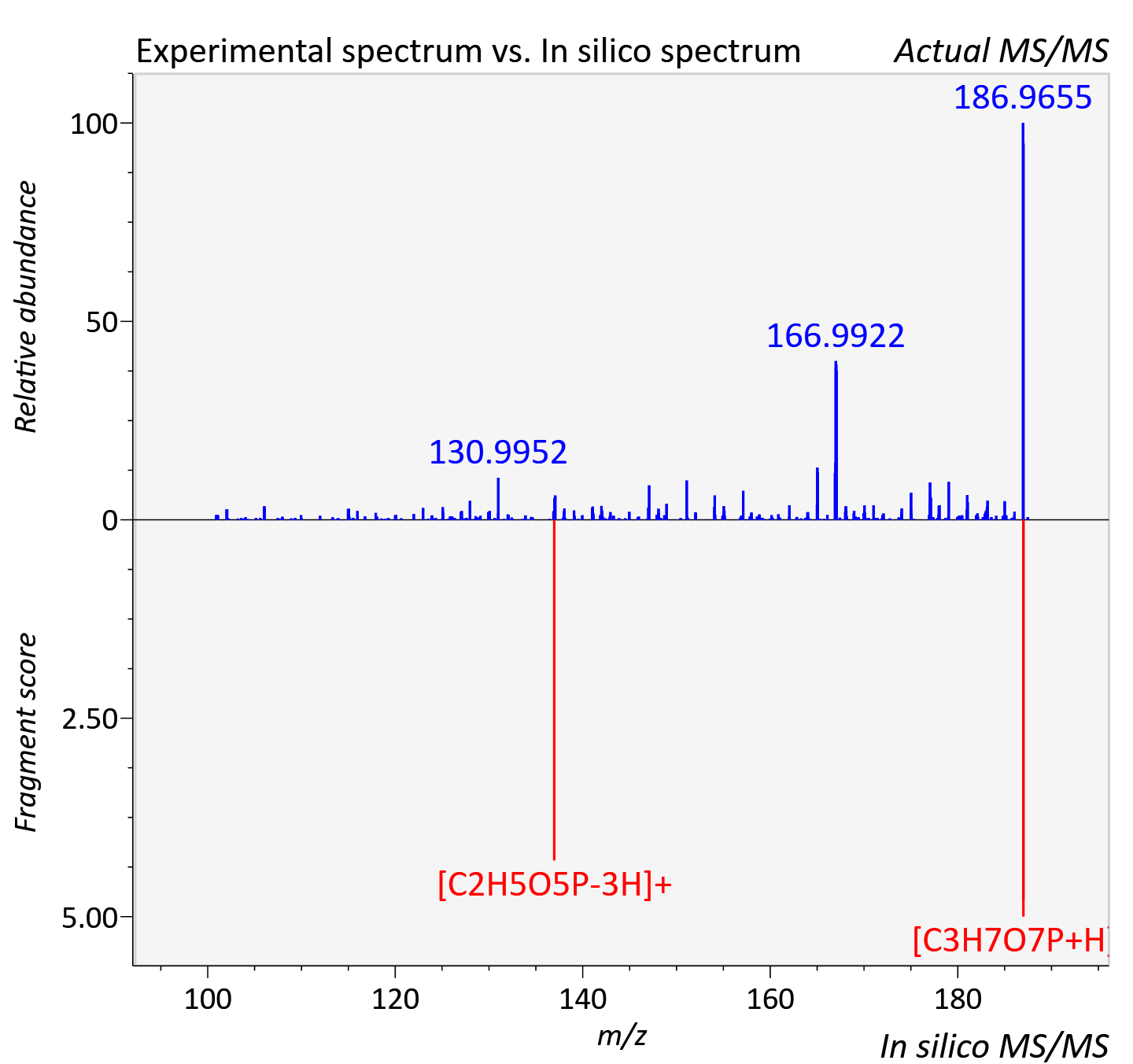
Experimental data
| Retention time | 16.84 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 187.002 | Theoretical mz | 187 |
| Error | 11.36 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.627 |
Identifiers and class information
| Inchi key | GXIURPTVHJPJLF-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(O)C(OP(=O)(O)O)CO |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| P12955 | PEPD | Xaa-Pro dipeptidase | T94823 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| P53602 | MVD | Diphosphomevalonate decarboxylase | T96862 | SEA |
