Compound details
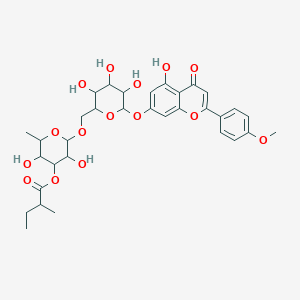
Acacetin 7-[3\'\'\'-(2-methylbutyryl)rutinoside]
| Compound ID | CDAMM00913 |
|---|---|
| Common name | Acacetin 7-[3\'\'\'-(2-methylbutyryl)rutinoside] | IUPAC name | [3,5-dihydroxy-2-methyl-6-[[3,4,5-trihydroxy-6-[5-hydroxy-2-(4-methoxyphenyl)-4-oxochromen-7-yl]oxyoxan-2-yl]methoxy]oxan-4-yl] 2-methylbutanoate |
| Molecular formula | C33H40O15 |
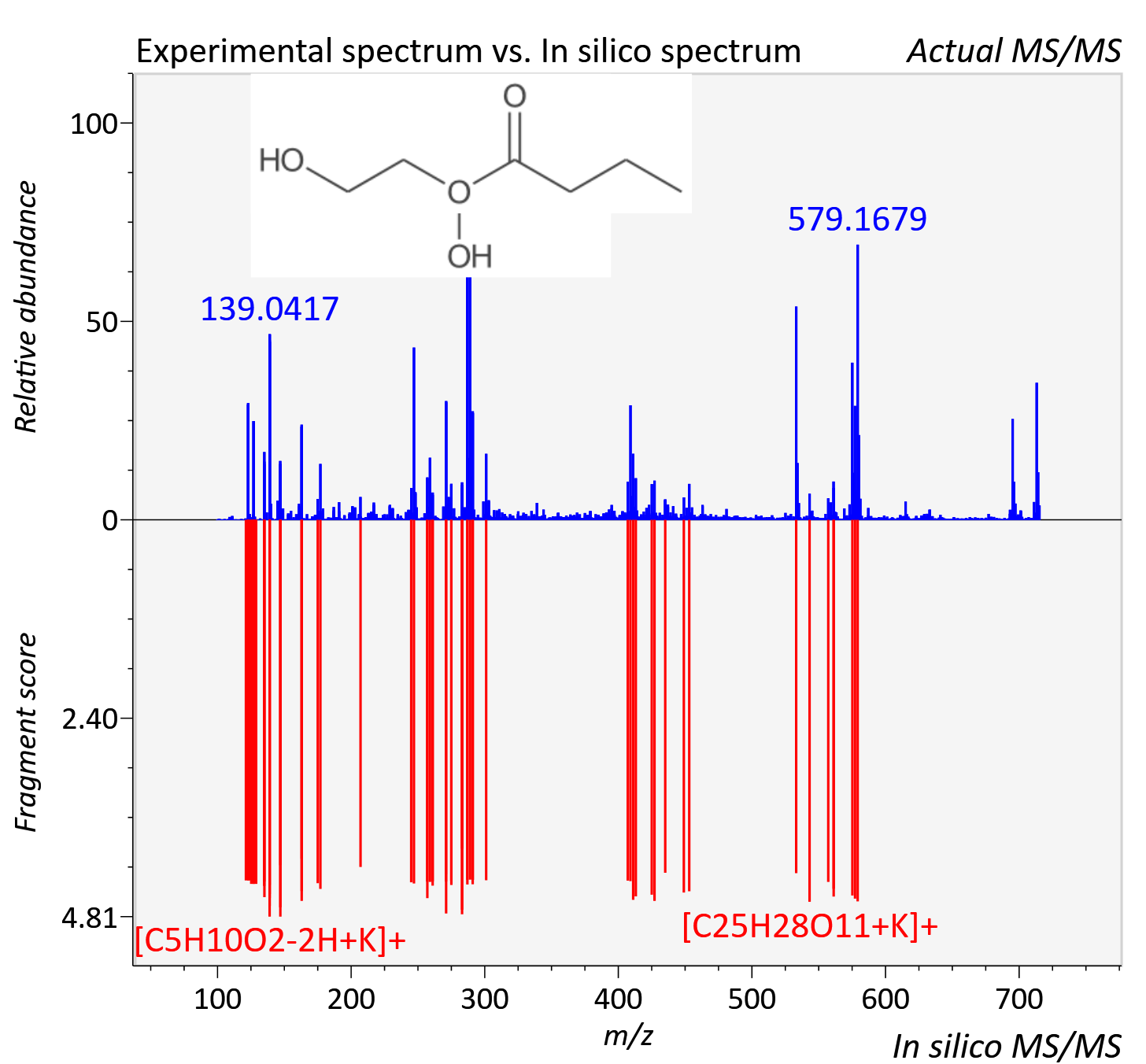
Experimental data
| Retention time | 0.27 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 715.202 | Theoretical mz | 715.2 |
| Error | 2.3 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.6718 |
Identifiers and class information
| Inchi key | QURQCINSSPAJEH-LEABYMNVNA-N |
|---|---|
| Smiles | O=C(OC1C(O)C(OCC2OC(OC3=CC(O)=C4C(=O)C=C(OC4=C3)C=5C=CC(OC)=CC5)C(O)C(O)C2O)OC(C)C1O)C(C)CC |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P15121 | AKR1B1 | Aldose reductase | T26623 | SwissTargetPrediction and SEA |
| P33527 | ABCC1 | Multidrug resistance-associated protein 1 | T11288 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P01375 | TNF | TNF-alpha | T20178 | SwissTargetPrediction |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| Q9NUW8 | TDP1 | Tyrosyl-DNA phosphodiesterase 1 | T33492 | SEA |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q9NPH5 | NOX4 | NADPH oxidase 4 | T29741 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q9NZ08 | ERAP1 | Endoplasmic reticulum aminopeptidase 1 | T72849 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
| P16220 | CREB1 | Cyclic AMP-responsive element-binding protein | T92098 | SEA |
| P30837 | ALDH1B1 | Acetaldehyde dehydrogenase | T99641 | SEA |
| Q9UK17 | KCND3 | Voltage-gated potassium channel Kv4.3 | T74500 | SEA |
| P68104 | EEF1A1 | Elongation factor 1A | T72269 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T26623 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P15121 | AKR1B1 |
| T26623 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P15121 | AKR1B1 |
| T11288 | DI0167 | Gout | [ICD-11: FA25] | P33527 | ABCC1 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T20178 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | P01375 | TNF |
| T20178 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | P01375 | TNF |
| T20178 | DI0351 | Psoriasis | [ICD-11: EA90] | P01375 | TNF |
| T20178 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P01375 | TNF |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T29741 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9NPH5 | NOX4 |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
| T99641 | DI0396 | Substance abuse | [ICD-11: 6C40] | P30837 | ALDH1B1 |
| T74500 | DI0004 | Acidosis | [ICD-11: 5C73] | Q9UK17 | KCND3 |
| T72269 | DI0252 | Melanoma | [ICD-11: 2C30] | P68104 | EEF1A1 |
