Compound details
(Z)-1-Methyl-2-(tridec-7-enyl)quinolin-4-one
| Compound ID | CDAMM00855 |
|---|---|
| Common name | (Z)-1-Methyl-2-(tridec-7-enyl)quinolin-4-one | IUPAC name | 1-methyl-2-tridec-7-enylquinolin-4-one |
| Molecular formula | C23H33NO |
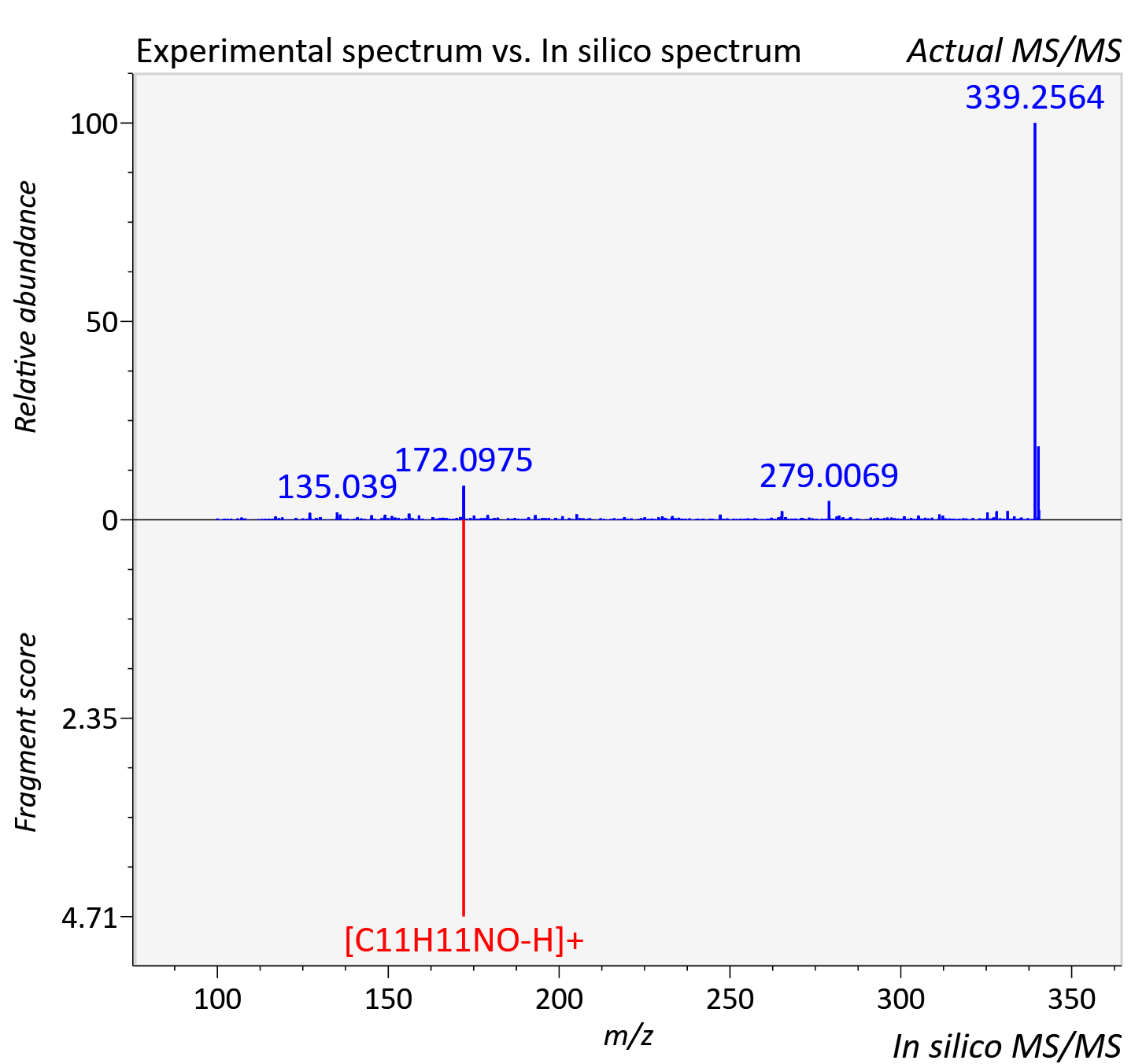
Experimental data
| Retention time | 20.21 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 340.259 | Theoretical mz | 340.263 |
| Error | 11.1 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2079 |
Identifiers and class information
| Inchi key | GXRKDSSARDBYHW-FPLPWBNLSA-N |
|---|---|
| Smiles | O=C1C=C(N(C=2C=CC=CC12)C)CCCCCCC=CCCCCC |
| Superclass | Organoheterocyclic compounds |
| Class | Quinolines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P09917 | ALOX5 | Arachidonate 5-lipoxygenase | T00140 | SEA |
| Q99685 | MGLL | Monoglyceride lipase | T18664 | SEA |
| P34972 | CNR2 | Cannabinoid receptor 2 | T37693 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| P35398 | RORA | Nuclear receptor ROR-alpha | T43206 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| Q15746 | MYLK | Myosin light chain kinase, smooth muscle | T97141 | SEA |
| P55786 | NPEPPS | Puromycin-sensitive aminopeptidase | T15610 | SEA |
| Q9NR97 | TLR8 | Toll-like receptor 8 | T48703 | SEA |
| Q9NYK1 | TLR7 | Toll-like receptor (TLR7/TLR9) | T46482 | SEA |
| Q9UGI6 | KCNN3 | Small conductance calcium-activated potassium channel protein 3 | T04388 | SEA |
| Q92753 | RORB | Nuclear receptor ROR-beta | T48812 | SEA |
| Q92887 | CMOAT | Multidrug resistance-associated protein 2 | T61792 | SEA |
| Q4ZFZ2 | FSHR | Follicle-stimulating hormone receptor | T68334 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T00140 | DI0037 | Asthma | [ICD-11: CA23] | P09917 | ALOX5 |
| T00140 | DI0147 | Filariasis | [ICD-11: 1F66] | P09917 | ALOX5 |
| T00140 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P09917 | ALOX5 |
| T18664 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q99685 | MGLL |
| T18664 | DI0409 | Tic disorder | [ICD-11: 8A05] | Q99685 | MGLL |
| T37693 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | P34972 | CNR2 |
| T37693 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P34972 | CNR2 |
| T43206 | DI0235 | Liver cancer | [ICD-11: 2C12] | P35398 | RORA |
| T43206 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P35398 | RORA |
| T97141 | DI0127 | Duodenal ulcer | [ICD-11: DA63] | Q15746 | MYLK |
| T15610 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P55786 | NPEPPS |
| T15610 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P55786 | NPEPPS |
| T48703 | DI0133 | Epidermal dysplasias | [ICD-11: EK90] | Q9NR97 | TLR8 |
| T48703 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | Q9NR97 | TLR8 |
| T48703 | DI0180 | Herpes simplex infection | [ICD-11: 1F00] | Q9NR97 | TLR8 |
| T48703 | DI0239 | Lupus erythematosus | [ICD-11: 4A40] | Q9NR97 | TLR8 |
| T48703 | DI0252 | Melanoma | [ICD-11: 2C30] | Q9NR97 | TLR8 |
| T48703 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | Q9NR97 | TLR8 |
| T48703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NR97 | TLR8 |
| T48703 | DI0393 | Squamous cell carcinoma | [ICD-11: 2B60-2D01] | Q9NR97 | TLR8 |
| T48703 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q9NR97 | TLR8 |
| T46482 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q9NYK1 | TLR7 |
| T46482 | DI0384 | Skin disease | [ICD-11: EA00-EM0Z] | Q9NYK1 | TLR7 |
| T04388 | DI0285 | Myelopathy | [ICD-11: 8B42] | Q9UGI6 | KCNN3 |
| T68334 | DI0144 | Female infertility | [ICD-11: GA31] | Q4ZFZ2 | FSHR |
