Compound details
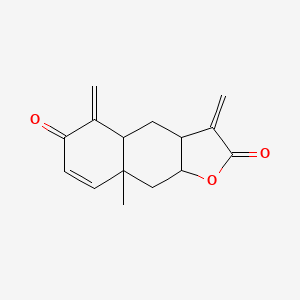
Encelin
| Compound ID | CDAMM00823 |
|---|---|
| Common name | Encelin | IUPAC name | 8a-methyl-3,5-dimethylidene-4,4a,9,9a-tetrahydro-3aH-benzo[f][1]benzofuran-2,6-dione |
| Molecular formula | C15H16O3 |
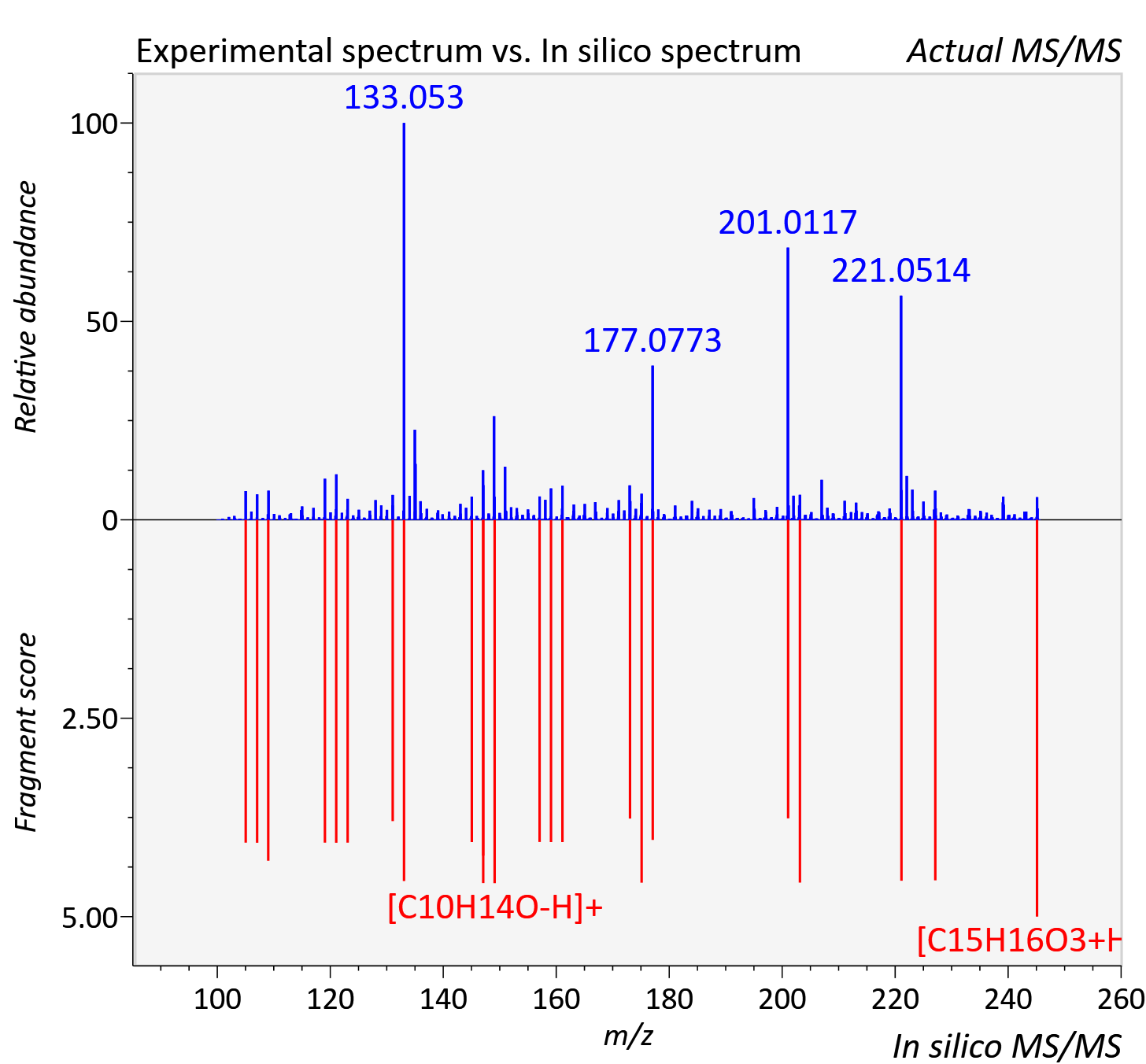
Experimental data
| Retention time | 15.59 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 245.118 | Theoretical mz | 245.117 |
| Error | 1.32 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.4247 |
Identifiers and class information
| Inchi key | LXMUZMFQJGRVFW-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC2CC3(C=CC(=O)C(=C)C3CC2C1=C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T66665 | DI0087 | Chronic pain | [ICD-11: MG30] | P35354 | PTGS2 |
| T66665 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P35354 | PTGS2 |
| T66665 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P35354 | PTGS2 |
| T66665 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P35354 | PTGS2 |
| T66665 | DI0227 | Knee osteoarthritis | [ICD-11: FA01] | P35354 | PTGS2 |
| T66665 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35354 | PTGS2 |
| T66665 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35354 | PTGS2 |
| T66665 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P35354 | PTGS2 |
| T66665 | DI0416 | Tuberculosis | [ICD-11: 1B10-1B12] | P35354 | PTGS2 |
