Compound details
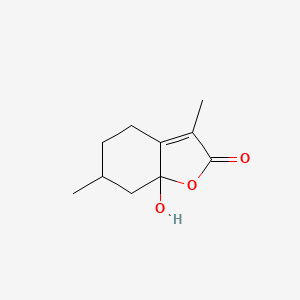
Peperinic acid
| Compound ID | CDAMM00812 |
|---|---|
| Common name | Peperinic acid | IUPAC name | 7a-hydroxy-3,6-dimethyl-4,5,6,7-tetrahydro-1-benzofuran-2-one |
| Molecular formula | C10H14O3 |
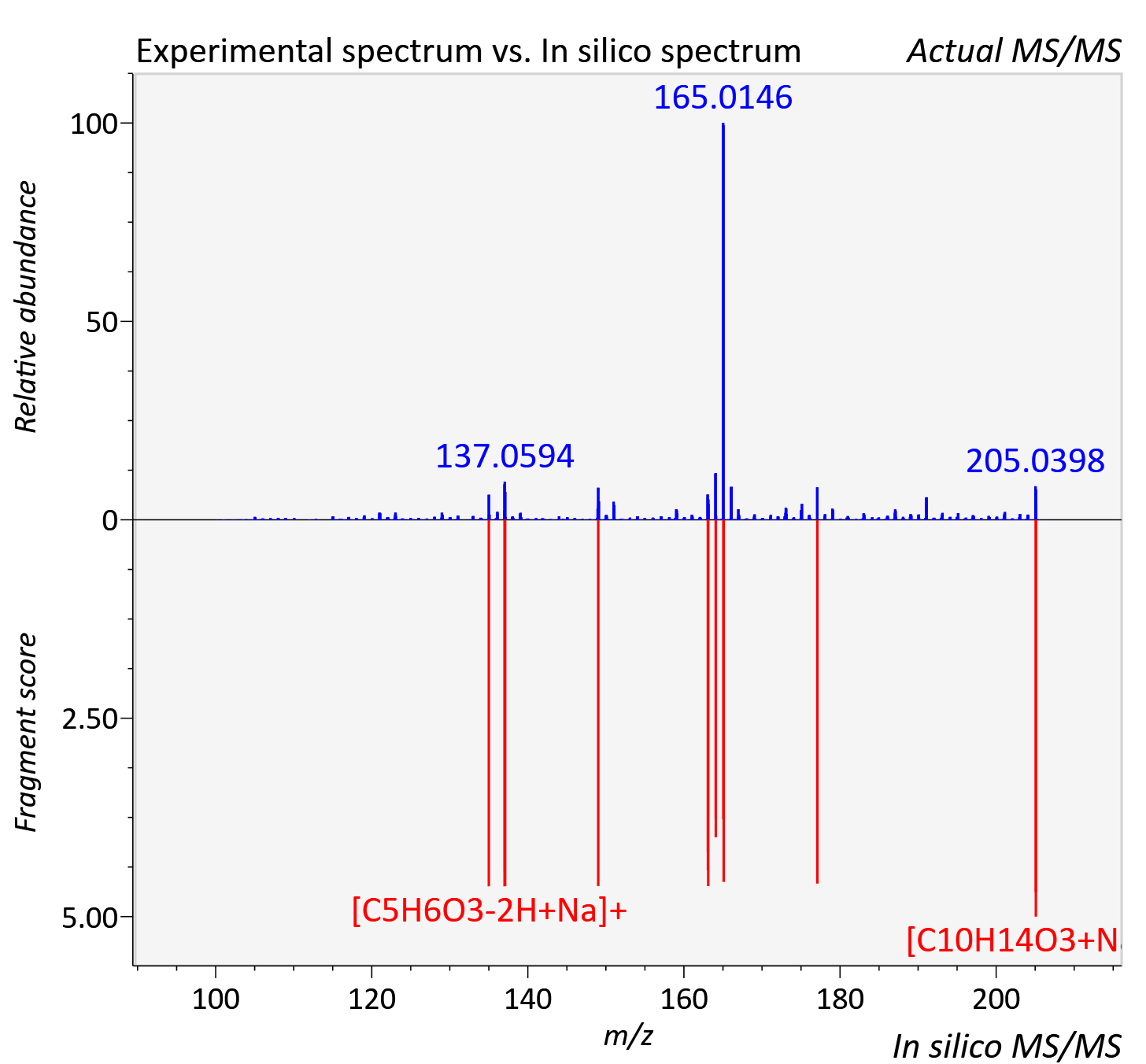
Experimental data
| Retention time | 18.2 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 205.086 | Theoretical mz | 205.083 |
| Error | 12.1 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.6806 |
Identifiers and class information
| Inchi key | LBNWZGLSMCTAQB-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC2(O)C(=C1C)CCC(C)C2 |
| Superclass | Organoheterocyclic compounds |
| Class | Benzofurans |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T31479 | DI0230 | Leishmaniasis | [ICD-11: 1F54] | P04054 | PLA2G1B |
| T31479 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P04054 | PLA2G1B |
| T31479 | DI0367 | Rosacea | [ICD-11: ED90] | P04054 | PLA2G1B |
| T31479 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P04054 | PLA2G1B |
