Compound details
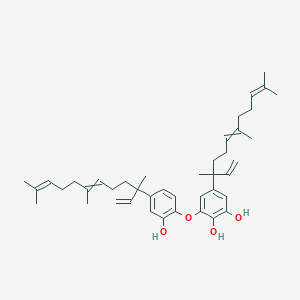
Peltatol C
| Compound ID | CDAMM00794 |
|---|---|
| Common name | Peltatol C | IUPAC name | 3-[2-hydroxy-4-(3,7,11-trimethyldodeca-1,6,10-trien-3-yl)phenoxy]-5-(3,7,11-trimethyldodeca-1,6,10-trien-3-yl)benzene-1,2-diol |
| Molecular formula | C42H58O4 |
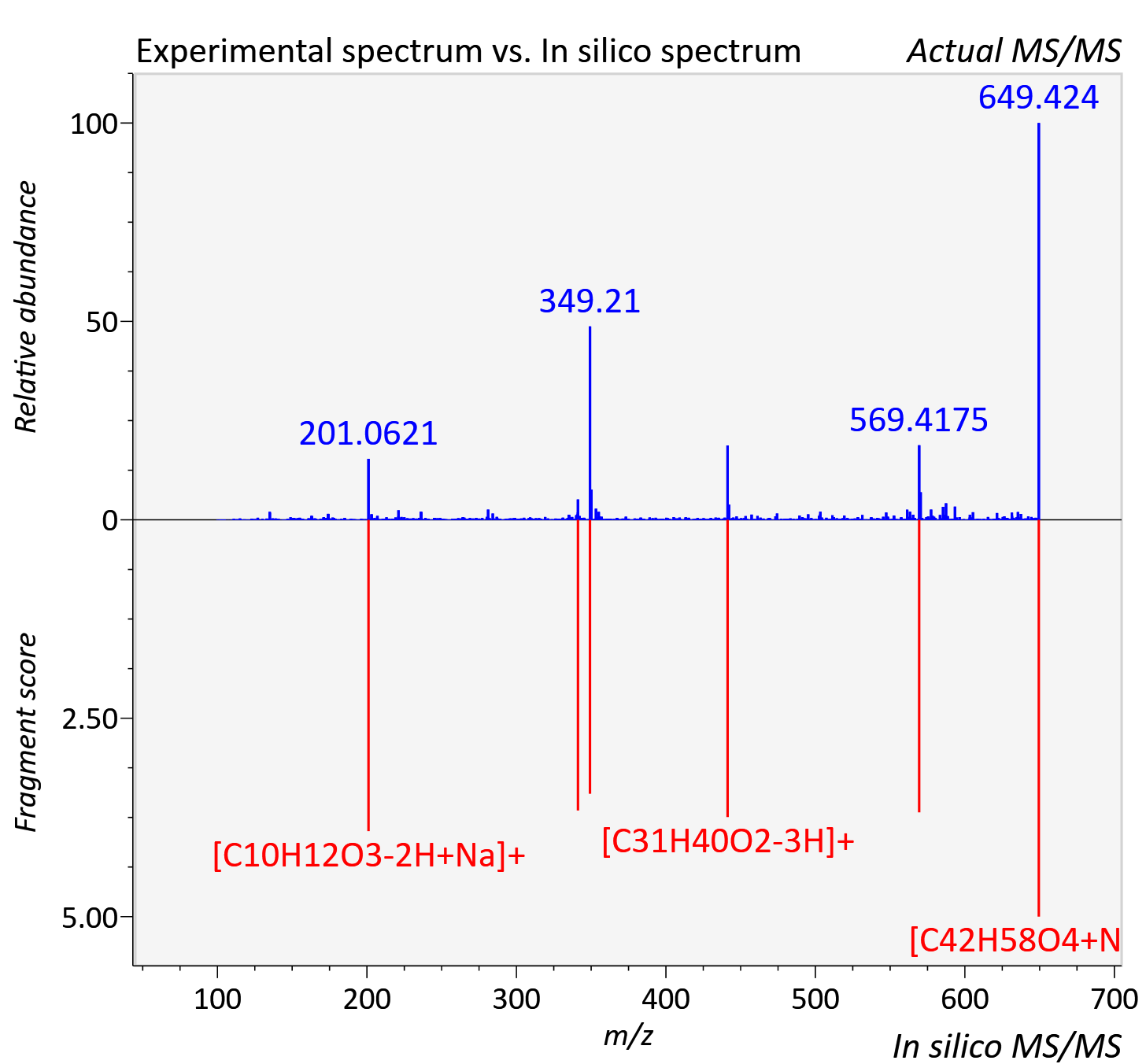
Experimental data
| Retention time | 15.34 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 649.424 | Theoretical mz | 649.423 |
| Error | 1.51 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.287 |
Identifiers and class information
| Inchi key | IIXXCYMHCAOUOF-RWHGWQAQNA-N |
|---|---|
| Smiles | OC=1C=C(C=C(OC2=CC=C(C=C2O)C(C=C)(C)CCC=C(C)CCC=C(C)C)C1O)C(C=C)(C)CCC=C(C)CCC=C(C)C |
| Superclass | Lignans, neolignans and related compounds |
| Class |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T93344 | DI0118 | Dermatophytosis | [ICD-11: 1F28] | Q14534 | SQLE |
| T93344 | DI0385 | Skin fungal infection disorder | [ICD-11: EA60] | Q14534 | SQLE |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
