Compound details
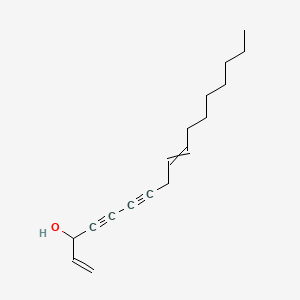
Panaxynol
| Compound ID | CDAMM00789 |
|---|---|
| Common name | Panaxynol | IUPAC name | heptadeca-1,9-dien-4,6-diyn-3-ol |
| Molecular formula | C17H24O |
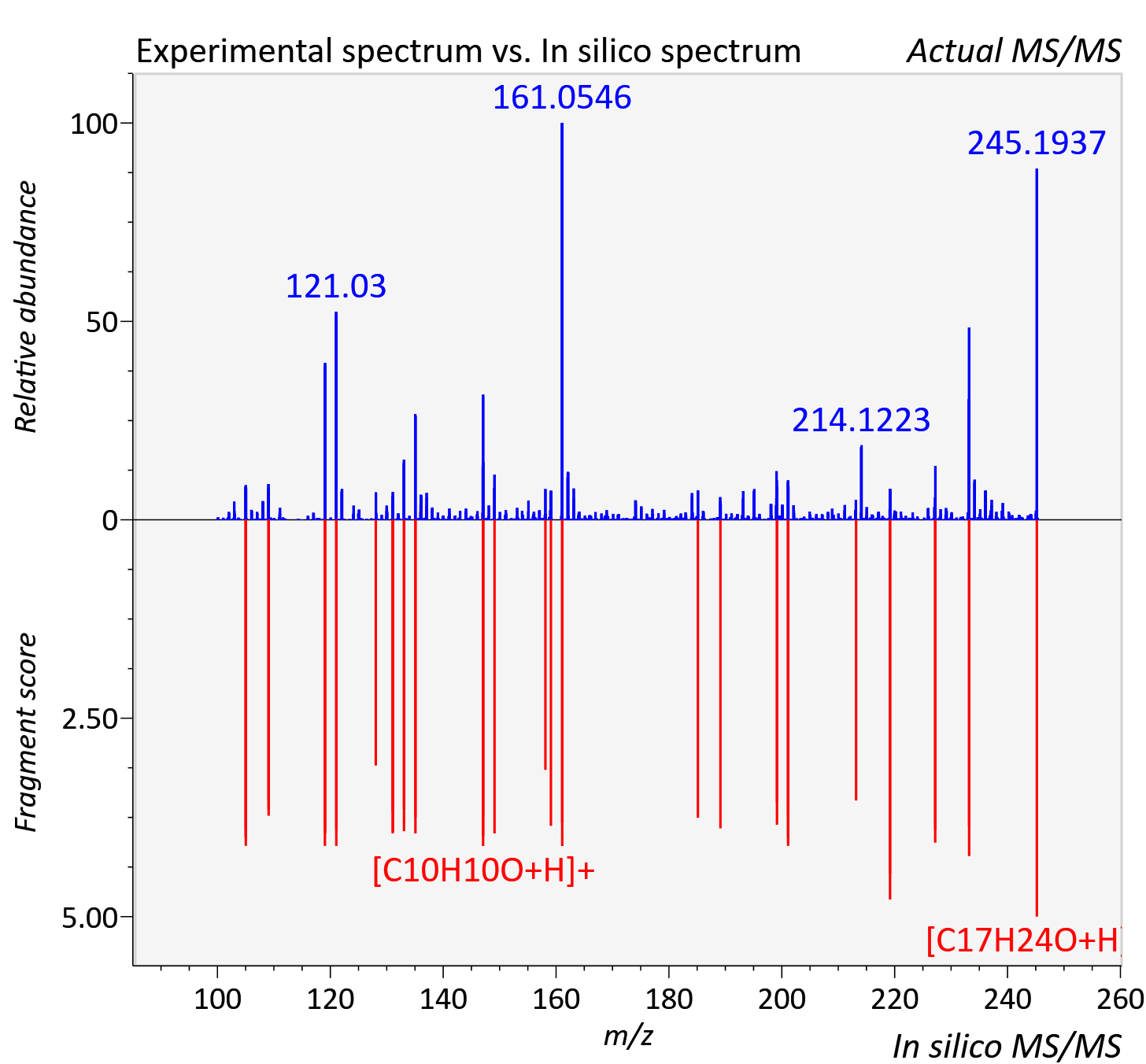
Experimental data
| Retention time | 16.19 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 245.192 | Theoretical mz | 245.19 |
| Error | 7.44 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2081 |
Identifiers and class information
| Inchi key | UGJAEDFOKNAMQD-ZHACJKMWNA-N |
|---|---|
| Smiles | OC(C#CC#CCC=CCCCCCCC)C=C |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P35372 | OPRM1 | Mu opioid receptor | T47768 | SwissTargetPrediction |
| P41145 | OPRK1 | Kappa Opioid receptor | T60693 | SwissTargetPrediction |
| P41143 | OPRD1 | Delta opioid receptor | T58992 | SwissTargetPrediction |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| Q9UP65 | PLA2G4C | Cytosolic phospholipase A2 gamma | T91113 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T47768 | DI0013 | Acute pain | [ICD-11: MG31] | P35372 | OPRM1 |
| T47768 | DI0059 | Bowel habit change | [ICD-11: ME05] | P35372 | OPRM1 |
| T47768 | DI0087 | Chronic pain | [ICD-11: MG30] | P35372 | OPRM1 |
| T47768 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P35372 | OPRM1 |
| T47768 | DI0105 | Cough | [ICD-11: MD12] | P35372 | OPRM1 |
| T47768 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P35372 | OPRM1 |
| T47768 | DI0124 | Digestive system disease | [ICD-11: DE2Z] | P35372 | OPRM1 |
| T47768 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P35372 | OPRM1 |
| T47768 | DI0228 | Large intestine motility disorder | [ICD-11: DB32] | P35372 | OPRM1 |
| T47768 | DI0317 | Opioid use disorder | [ICD-11: 6C43] | P35372 | OPRM1 |
| T47768 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35372 | OPRM1 |
| T47768 | DI0349 | Pruritus | [ICD-11: EC90] | P35372 | OPRM1 |
| T47768 | DI0374 | Sensation disturbance | [ICD-11: MB40] | P35372 | OPRM1 |
| T60693 | DI0304 | Non-specific cutaneous vascular symptom | [ICD-11: ME64] | P41145 | OPRK1 |
| T60693 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P41145 | OPRK1 |
| T60693 | DI0349 | Pruritus | [ICD-11: EC90] | P41145 | OPRK1 |
| T58992 | DI0059 | Bowel habit change | [ICD-11: ME05] | P41143 | OPRD1 |
| T58992 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P41143 | OPRD1 |
| T58992 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P41143 | OPRD1 |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
